Caires, Lyra, Hentschke, da Silva, de Araújo, Sant’Anna, and Nunes: Polyphasic delimitation of a filamentous marine genus, Capillus gen. nov. (Cyanobacteria, Oscillatoriaceae) with the description of two Brazilian species
ABSTRACT
Lyngbya C. Agardh ex Gomont is a nonheterocytous cyanobacterial genus whose evolutionary history is still poorly known. The traditionally defined Lyngbya has been demonstrated to be polyphyletic, including at least five distinct clades, some of which have been proposed as new genera. Intraspecific diversity is also clearly underestimated in Lyngbya due to the lack of unique morphological characters to differentiate species. In this study, we describe the new genus Capillus T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes from benthic marine environments, including two new Brazilian species (here described as C. salinus T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes, and C. tropicalis T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes), and two species yet to be described, one of them from India (Capillus sp. 2.1), and the other from United States of America, based on strain PCC 7419. Capillus species presented cross-wise diagonal fragmentation, assisted or not by necridic cells, which has not been previously mentioned for Lyngbya. Ultrastructural analyses showed that C. salinus and C. tropicalis have numerous gas vesicles, which are rarely described for benthic marine species. The new genus formed a well-supported clade, and the D1-D1′ and Box B secondary structures of internal transcribed spacer also supported the proposal of its new species. These findings help to clarify the diversity of species in the Lyngbya complex and the taxonomy of the group, and highlight the need of further floristic surveys in tropical coastal environments, which remain poorly studied.
Key words: Brazil; Cyanobacteria; diversity; new genus; polyphasic evaluation
Abbreviations
ITS
internal transcribed spacer
16S rRNA
small subunit of prokaryotic ribosome gene
INTRODUCTION
Lyngbya C. Agardh ex Gomont (Oscillatoriaceae) is a nonheterocytous cyanobacterial genus whose evolutionary history is still poorly known ( Komárek and Anagnostidis 2005). Numerous studies have demonstrated that the traditionally defined Lyngbya is polyphyletic and includes at least six distinct genetic clades ( Sharp et al. 2009, Engene et al. 2010, 2013, Komárek et al. 2013, McGregor and Sendall 2014, Stoyanov et al. 2014, Caires et al. 2018). Consequently, Lyngbya, as currently intended, is a polyphyletic group, probably including several cryptogenera (i.e., lineages that cannot be distinguished by morphology and / or ecology, but only using molecular methods), comprising well-delimitated clades, which lack morphological exclusive characters ( Komárek 2016).
Recently, five of the clades traditionally identified as Lyngbya have been proposed as new genera, three including marine species [ Moorea N. Engene, E. C. Rottacker, J. Kaštovský, T. Byrum, H. Choi, M. H. Ellisman, J. Komárek and W. H. Gerwick ( Engene et al. 2012), Okeania N. Engene, V. J. Paul, T. Byrum, W. H. Gerwick, A. Thor and M. H. Ellisman ( Engene et al. 2013), and Neolyngbya T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes ( Caires et al. 2018)]; and two including freshwater species [ Limnoraphis J. Komárek, E. Zapomělová, J. Šmarda, J. Kopecký, E. Rejmánková, J. Woodhouse, B. A. Neilan and J. Komárková ( Komárek et al. 2013) and Microseira G. B. McGregor and B. C. Sendall ( McGregor and Sendall 2014)]. Species delimitation is challenging in this group of Cyanobacteria, especially as most taxonomic studies are restricted to morphological characters, which frequently overlap among species ( Komárek et al. 2014). Therefore, the diversity of species in the Lyngbya complex is underestimated. Komárek and Anagnostidis (2005) referred only 11 marine morphospecies as Lyngbya in their review: L. aestuarii Liebman ex Gomont, L. agardhii Gomont, L. atrovirens Harvey, L. bouillonii Hoffmann et Demoulin, L. confervoides C. Agardh ex Gomont (type species of the genus), L. majuscula Harvey ex Gomont, L. meneghiniana Gomont, L. salina Kützing ex Gomont, L. semiplena J. Agardh ex Gomont, L. sordida Gomont, and L. usteri Schmidle. Brazil presents a long coast where the presence of several cyanobacteria species has been recorded. Despite the broad recognition of the importance of polyphasic characterization for studying cyanobacteria, only the study carried out by Caires et al. (2018) in Brazil used this approach for analyzing Lyngbya populations. The few studies performed for this cyanobacterial genus with Brazilian sampling have examined the diversity through morphological taxonomy and ecological features, such as Sant’Anna et al. (1985), Baeta-Neves (1991), Sant’Anna (1995, 1997), Crispino and Sant’Anna (2006), and Caires et al. (2013). Five Lyngbya species were referred to marine environments of Brazil: Lyngbya aestuarii Liebman ex Gomont, L. confervoides C. Agardh ex Gomont, L. majuscula Harvey ex Gomont, L. semiplena J. Agardh ex Gomont, and L. sordida Gomont ex Gomont ( Werner et al. 2017). Engene et al. (2012) synonymized L. majuscula and L. sordida under Moorea producta N. Engene, E. C. Rottacker, J. Kaštovský, T. Byrum, H. Choi, M. H. Ellisman, J. Komárek and W. H. Gerwick. However, Brazilian species above mentioned need to be reevaluated using a combination of methods, including molecular tools, to confirm their taxonomic status. The presence of cryptogenera and species complexes in Oscillatoriaceae implies the necessity of further investigating this group using combined tools to show the real biodiversity of the Lyngbya complex, especially in undersampled areas such as the Brazilian coast. Here we use a polyphasic approach, which is based on morphological, molecular, ultrastructural, and ecological data. Thus, we contribute to the knowledge on the taxonomy of this challenging group, describing the new marine genus Capillus which was dismembered from Lyngbya complex. This new genus is represented by two new Brazilian species (Capillus salinus sp. nov. and Capillus tropicalis sp. nov.), besides two species yet to be described, one of them from India, and the other from USA.
MATERIALS AND METHODS
Sampling and cultivation of cyanobacteria
We sampled populations on the tropical Brazilian coast, at Pedra do Sal Beach, in Salvador municipality (Bahia State) (12°57′06″ S, 38°20′42″ W), and at Arembepe Beach, in Camaçari municipality (Bahia State) (12°44′27″ S, 38°09′00″ W) ( Fig. 1). Sampling took place in the supralittoral zone, during low spring tides. We used metal spatulas for collecting epilithic samples and a refractometer (model 211; Biobrix, São Paulo, Brazil) for measuring salinity. We transported the material in seawater and once in the laboratory, samples were inoculated in SWBG-11 liquid medium ( Rippka et al. 1979). Vitamin B 12 (2 mL L −1 from a stock solution of 1 mg mL −1; Sigma-Aldrich, St. Louis, MO, USA) was added to stimulate growth. Afterwards, the specimens were isolated by selecting single trichomes using pipette Pasteur with hook-like modified tips. The specimens were grown under a 14 : 10 h (light : dark) cycle under constant irradiation at 30 μmol photons m −2 s −1 (white fluorescent lamps) and at 25 ± 1°C ( Jacinavicius et al. 2012). All isolated strains were maintained in the Culture Collection associated with the herbarium ALCB of the Biology Institute from Universidade Federal da Bahia (UFBA), Brazil. Subsamples from all strains were also preserved in formaldehyde (4%) and deposited in the mentioned herbarium.
Morphological and ultrastructural evaluation
We studied the strains using a light microscope equipped with a QImaging GO-3 digital camera (Olympus CX31RTS5, Tokyo, Japan). A minimum of 20 filaments from natural populations and also from each culture were measured using the AxioVs40 v4.8.2.0 imaging system (Carl Zeiss, Jena, Germany).
For transmission electron microscopy, the cyanobacterial filaments from natural populations and from cultures were fixed overnight in modified Karnovsky (final concentrations: 2% glutaraldehyde, 0.05 M sodium cacodylate-pH 7.2, 0.001 M calcium chloride, and 2% paraformaldehyde) ( Karnovsky 1965). Biomass was washed three times with 0.01 M sodium cacodylate buffer and post-fixed with 1% osmium tetroxide in the same buffer at room temperature for 2 h. The biomass remained in 2.5% uranyl acetate overnight at 4°C after another wash in the same buffer. Subsequently, the filaments were dehydrated in an acetone series (30, 50, and 70% for 5 min each and 90 and 100% for 20 min each) and embedded in Poly/Bed 812 Embedding Kit resin (Polysciences, Warrington, PA, USA). We cut ultrathin sections using an ultramicrotome (Leica, Mannheim, Germany); these sections were post-stained with 2.5% uranyl acetate and lead citrate. The obtained sections were analyzed with a JEOL 1230 transmission electron microscope (JEOL, Tokyo, Japan). Strains were also processed for scanning electron microscopy. We fixed biomass samples overnight in modified Karnovsky at room temperature. Subsequently, the samples were washed three times in 0.01 M sodium cacodylate buffer and were dehydrated in an alcohol series (25, 50, 70, 90, and 100% for 15 min each) followed by critical point drying (Leica CPD030). We placed the specimens on stubs, coated them with sputtered gold (Denton Vacuum Desk IV, Moorestown, NJ, USA) and analyzed each specimen using a JEOL 6390LV scanning electron microscope (JEOL).
Molecular analyses
Specimens were macerated in liquid nitrogen to extract genomic DNA according to Fiore et al. (2000). Almost complete16S rRNA gene and the 16S–23S internal transcribed spacer (ITS) regions were amplified by polymerase chain reaction (PCR) using the primers 27F1 ( Neilan et al. 1997) and 23S30R ( Taton et al. 2003) in a Veriti 96-Well Thermal Cycler (Applied Biosystems, Foster City, CA, USA). For PCR reactions, we used 10 ng of genomic DNA, 0.5 μM of each primer, 200 μM of dNTPs, 2.0 mM of MgCl 2, l × PCR buffer, and 1.5 U Taq DNA polymerase (Invitrogen, São Paulo, Brazil); the total volume was 25 μL. Molecular markers were amplified under the following conditions: one cycle of 5 min at 94°C; 10 cycles of 45 s at 94°C, 45 s at 57°C, and 2 min at 72°C; 25 cycles of 45 s at 94°C, 45 s at 54°C, and 2 min at 72°C; the final elongation step of 7 min at 72°C. PCR products were analyzed by agarose gels (1% w/v) using the Low DNA mass ladder (Invitrogen, Carlsbad, CA, USA). PCR products were cloned using the pGEM-T Easy vector system (Promega, Madison, WI, USA) according to the manufacturer’s protocol. Chemo-competent Escherichia coli DH5α cells were transformed and plated for blue-white colony screening ( Sambrook and Russell 2001). The plasmids containing gene fragments were extracted by alkaline lysis method ( Birnboim and Doly 1979). We sequenced plasmid DNA with the plasmid primers M13F/M13R and the 16S rRNA with the following internal primer sets to cover the entire fragment: 357F/357R, 704F/704R, and 1114F/1114R ( Lane 1991). Genetic sequencing was done at Instituto Gonçalo Muniz (FIOCRUZ–Bahia State, Brazil).
Phylogenetic and secondary-structures analyses
Geneious v6.0.6 ( Drummond et al. 2011) was used to assemble and edit the sequences. Our dataset to compose the phylogenetic analyses comprised sequences that were identified by BLAST search as closely related to sequences obtained in our study, besides sequences representing the Oscillatoriaceae genera which were included in this dataset too (according to new classification proposed by Komárek et al. 2014). We used a total of 95 OTUs (operational taxonomic unit) available at GenBank ( http://www.ncbi.nlm.nih.gov/GenBank/; searched on February 2017) ( Supplementary Table S1). Additionally, Nostoc punctiforme (GenBank accession No. GQ287652) was included as the outgroup. The sequences obtained in this study were deposited in GenBank under the numbers KY824052 and MF190468. The evolutionary model (GTR + I + G) was identified by MrModeltest 2.3 ( Nylander 2008) using the Akaike Information Criterion ( Posada and Buckley 2004). Reconstruction of phylogenetic trees by the maximum-likelihood algorithm (ML) ( Stamatakis et al. 2008) was carried out with RaxML 8.2.3 ( Stamatakis 2006), which was tested using 1,000 bootstrapping replicates. Bayesian Inference analyses ( Mau et al. 1999) were conducted with MrBayes 3.1 software ( Ronquist et al. 2012), in two independent runs, with four chains each, for 5 × 10 6 Markov Chain Monte Carlo generations, which were verified to confirm the consistency of stationary-phase likelihood in our runs. Results from these phylogenetic analyses are expressed in bootstrap percentage (BP) for ML, and posterior probability (PP) for Bayesian analyses. Values lower than 70%, for both results, are not shown. The construction of secondary structures of the 16S–23S rRNA ITS sequences was determined using Mfold WebServer ver. 2.3, which was set at the default parameters ( Zuker 2003). The differently conserved and variable regions were identified according to Iteman et al. (2002). Previous analysis has shown that our sequences do not have tRNA Ile and tRNA Ala.
RESULTS
The new marine genus Capillus is represented by two new Brazilian species (C. salinus sp. nov. and C. tropicalis sp. nov.), and two species yet to be described, one of them from India (Capillus sp. 2.1), and the other from USA, based on strain PCC 7419 (accession numbers from GenBank: NR114680, AJ000714, NR112110, and AB039013).
Morphological evaluation and ultrastructure
Capillus T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes gen. nov
Thallus forming extensive fasciculate mats or small clusters, brown, brown-green, olive-green to dark-green, dark blue-green to black. Filaments straight, isopolar, unbranched, (8-) 12–30 (−46) μm diam. Sheaths hyaline or bright yellow to yellow-brown, firm, thin or thick, sometimes lamellated. Trichomes not or slightly constricted at the cross-walls, not or slightly attenuated towards ends, (6.6-) 10–21 (−24) μm diam., sometimes some trichomes become contorted to originate the necridial regions. Cells 1.4–5.6 (−6) μm long, 1.2–11 times wider than long. Cross-walls often granulated, sometimes with large granules. Apical cells rounded, conical, conical-rounded, flat-rounded, truncate, rarely capitate, sometimes thickened, without calyptra. Hormogonia formed by fragmentation perpendicular to the longitudinal axis of the trichome helped by necridic cells, or in diagonal position, in this case helped or not by necridic cells.
Etymology: Reference to the macroscopic aspect of the thallus, which has a hair-clusters-like shape. Diagnosis: Genus delimited by molecular and phylogenetic analyses based on the 16S rRNA gene and 16S–23S ITS region with distinctive D1-D1′ structures. Type species: Capillus salinus T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes sp. nov.
We summarized a morphological comparison among Capillus gen. nov. and the closely related genera Limnoraphis and Lyngbya ( Table 1).
Capillus salinus T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes sp. nov. (Fig. 2)
Thallus fasciculate, forming extensive mats, brown. Filaments straight, 14–16.4 μm diam. Sheaths hyaline, thin or slight thick, firm. Trichomes cylindrical, slightly constricted at the cross-walls, and sometimes slightly attenuated towards ends, 12.1–14.6 μm diam. Discoid cells, 1.7–2.5 μm long, 5.8–7 times wider than long. Cell content dark-green, granulated. Cross-walls granulated, sometimes with large granules. Apical cells rounded, conical-rounded, conical, flat-rounded, without thickening. Hormogonia formed by cross-wise diagonal fragmentation of the trichome, helped or not by necridic cells.
Ultrastructural notes: Thylakoidal membranes presented parietal or irregular arrangement, which are distributed in different regions of the cell; cyanophycin granules occurred frequently near the cross-walls. Gas vesicles were observed usually in the inter-thylakoid space, frequently forming irregular clusters (aerotopes) near the cell wall and cross-wall. Diagnosis: Thallus fasciculate, brown, forming wide clusters, and delimited by molecular and phylogenetic analyses based on the 16S rRNA gene and 16S–23S ITS region with distinctive D1-D1′ and Box B structures. Etymology: Named after the high salinity at the collection site. Holotype: Brazil, Bahia State, Pedra do Sal Beach, Salvador city, 12°57′06″ S, 38°20′42″ W, Coll. T. A. Caires and E. S. T. Pina 842, Oct 12, 2015 (ALCB 114379). Paratype: Coll. J. M. C. Nunes E10, Nov 22, 2011 (ALCB 103416). Habit and habitat: Epilithic in tidal pools with high salinity (50 psu) in the supralittoral zone. This species formed large mats surrounding the full extent of the tidal pool.
Capillus tropicalis T. A. Caires, C. L. Sant’Anna et J. M. C. Nunes sp. nov. (Fig. 3)
Thallus fasciculate, forming small clusters, black. Filaments straight, 12–15.8 μm diam. Sheaths hyaline, thick, firm, frequently lamellated. Trichomes slightly constricted at the cross-walls, 10.1–12.9 μm diam., someone occasionally becomes contorted to originate the necridial regions. Cells 1.4–2.1 μm long, 1.2 times wider than long. Cell content green, sometimes granulated, and with numerous gas vesicles. Cross-walls granulated. Apical cell rounded or conical-rounded, rarely capitate. Hormogonia formed by straight fragmentation helped by necridic cells, or in diagonal position, in this case helped or not by necridic cells.
Ultrastructural notes: This species presented thylakoidal membranes with irregular arrangement and numerous gas vesicles occurring in great number near both cross and cell walls. Holotype: Brazil, Bahia State, Camaçari city, Arembepe Beach, 12°44′27″ S, 38°09′00″ W, Coll. T. A. Caires 794, Jan 21, 2015 (ALCB 114392). Type strain: Capillus tropicalis ALCB 114392 (GenBank: MF190468). Etymology: Occurrence in Brazilian tropical marine environments. Diagnosis: Thallus fasciculate, forming small clusters, black, and delimited by molecular and phylogenetic analyses based on the 16S rRNA gene and 16S–23S ITS region with distinctive D1-D1′ and Box B structures. Habit and habitat: Epilithic in protected reef region in mediolittoral zone.
Capillus sp. (strain PCC 7419; accession numbers from GenBank: NR114680, AJ000714, NR112110, and AB039013)
Morphological and ultrastructural notes: The strain 7419 from the Pasteur Culture Collection of Cyanobacteria (PCC), have been analyzed by many authors, like Porta and Hernández-Mariné (2005), who described some morphological and ultrastructural features to this strain. According to these authors, strain PCC 7419 has the following features: trichomes slightly constricted at the cross-walls, 8.1–9.7 μm diam., rounded apical cells, multilayered sheath; glycogen and cyanophycin granules, and carboxysomes present in the cell content; fragmentation of trichomes is helped by necridic cells. Regarding to ultrastructural features, strain PCC 7419 presents parietal thylakoid arrangement, and gas vesicles and aerotopes were not observed by Porta and Hernandéz-Mariné (2005).
The strain PCC 7419 is usually considered as type strain of the species L. aestuarii Liebman ex Gomont, since, according to Castenholz et al. (2001), it appears to be equivalent to the morphotype L. aestuarii, and it is structurally very similar to large species of Oscillatoria. As we did not have the opportunity to analyze this strain morphologically and ultrastructurally, we have treated this strain only as Capillus sp., and no as a new combination “ Capillus aestuarii.” The specific epithet to Capillus sp. based on the strain PCC 7419, can be proposed after a detailed morphological and ultrastructural study.
Capillus sp. (strain 2.1; accession number from GenBank: KF644563)
Morphological and ultrastructural notes: Mukherjee et al. (2015) provides only ecological features and one image with scale to strain Capillus sp. 2.1. From this image, it is possible to observe that the trichomes of this species present discoid cells and cell diam. ranging from 7.5 to 8.5 μm. As we have suggested to Capillus sp., the specific epithet to Capillus sp. 2.1 can be defined after more morphological and ultrastructural studies. Habit and habitat: Strain isolated from a bank of a river with highly saline water in India.
Phylogenetic and secondary structures analyses
We generated partial sequences of 16S rRNA (1.274–1.477 bp), which were aligned with a total of 95 sequences of Oscillatoriaceae for phylogenetic analyses. Our analyses corroborated the polyphyly of the traditionally defined Lyngbya genus. Capillus gen. nov. formed a well-supported clade (100% PP and 70% BP), which was sister taxon to the genus Limnoraphis ( Fig. 4). Our analyses included also one sequence from India ( Lyngbya sp. KF644563), and four sequences of strains previously identified as L. aestuarii, which were produced from the same strain (PCC 7419) by different studies. The Limnoraphis clade (76% BP and 98% PP), and the L. confervoides clade (100% BP and 100% PP), which is the type species of Lyngbya ( Gomont 1892), were also well-supported. Five clades, which were well-supported [ Lyngbya (?) clades: I, II, III, IV, and V] ( Fig. 4), containing sequences designated as Lyngbya were identified in our phylogeny. Besides, four sequences also tagged as Lyngbya (underline sequences) ( Fig. 4), were spread along our phylogenetic reconstruction. Considering the lack of both tRNA Ile and tRNA Ala in our sequences, we analyzed three conserved regions (D1-D1′, Box B, and V3) in the 16S–23S rRNA ITS sequences (565 to 595 bp) of C. salinus sp. nov. and C. tropicalis sp. nov. The secondary structures for each region are shown in Fig. 5. The D1-D1′ domain presented 61 and 58 nucleotides for C. salinus sp. nov. and C. tropicalis sp. nov., respectively. The related taxa Limnoraphis sp. (JX006094 and JX006095) presented the D1-D1′ domain with 58 nucleotides. Regarding the D1-D1′ helix, C. salinus sp. nov. presents a basal stem with 6 bp and no free nucleotides opposing the first lateral bulge. In C. tropicalis and Limnoraphis, the basal stem is shorter because of nucleotides substitutions in this region, leading to the presence of a 5 residue right bulge for C. salinum and a 7 residue asymmetrical internal loop for C. tropicalis. Limnoraphis has an 8 residue asymmetrical internal loop (nucleotides colored in Fig. 5). The medial stem of Capillus and Limnoraphis presents a single base left bulge (red box in Fig. 5) in different positions of the helices. The terminal loop was also variable in sequences and lengths among these taxa. The Box B helices were very different between the new species, presenting 34 nucleotides in C. salinus and 24 nucleotides in C. tropicalis. The V3 helices of Capillus were very similar, differing only by a nucleotide substitution in the medial stem (colored in Fig. 5) and in the deletion of one adenine in the terminal loop of C. tropicalis.
DISCUSSION
Morphological and ultrastructural analyses
The analyzed populations presented several morphological and ecological features similar to the Lyngbya complex, such as: the aspect of the thallus, cylindrical trichomes, discoid cells, and sheaths obligatorily surrounding only one trichome. Hence, it is difficult to discern Capillus gen. nov. from Lyngbya using only morphological characters. Thus, Lyngbya is considered a polyphyletic group, probably including many cryptogenera ( Komárek 2016). However, accurate molecular and ultrastructural analyses revealed that species of Capillus are not closely related to Lyngbya confervoides clade, which is considered the type species of this genus. The kind of fragmentation of the trichome for the formation of the hormogonia is a feature that can help in the distinction between these two genera. Trichome fragmentation due to the formation of necridial cells, which cause breaking points perpendicular to the longitudinal axis of the trichome, is typical of the genus Lyngbya ( Komárek and Anagnostidis 2005). Capillus Brazilian species also presented diagonal fragmentation (assisted or not by necridic cells), which has not been previously reported for Lyngbya. Ultrastructural analyses showed that C. salinus sp. nov. and C. tropicalis have irregular arrangement of thylakoids, as described for Oscillatoriaceae ( Komárek et al. 2014), while Porta and Hernandéz-Mariné (2005) have mentioned parietal thylakoids arrangement to Capillus sp. (strain PCC 7419). Our analyses also showed that the Brazilian species have numerous gas vesicles in the inter-thylakoid space and a pronounced accumulation of these structures in specific regions of the cell. According to Komárek and Anagnostidis (2005), aerotopes are reported from Lyngbya only rarely and in planktic species, but not in benthic species. Capillus sp. 2.1, not morphologically described in this study, has not yet been analyzed with ultrastructural studies. Porta and Hernandéz-Mariné (2005) carried out the ultrastructural study on strain PCC 7419, and gas vesicles are not reported. The gas vesicles are environmentally conditioned according to Komárek (2018), and the strain PCC 7419 has been maintained on culture for long time, which may have influenced the occurrence of these intracellular structures. Thus, it is not possible to confirm if this strain presented gas vesicles in its cell content. The occurrence of gas vesicles in benthic cyanobacterial species is quite interesting. This feature can be associated with the environmental dispersion of hormogonia, which can therefore remain in the water column and reach places relatively distant from their original location. Komárek (2016) stated that in some species the gas vesicles are observed only occasionally, as in hormogonia. However, in C. salinus and C. tropicalis these structures occurred frequently not only in hormogonia, but also in the whole adult filament. So, further studies with ecological and physiological approaches are necessary to better understand the importance of these structures in these benthic organisms.
Phylogenetic analyses and ITS secondary structures
It has been demonstrated that the traditionally defined Lyngbya is polyphyletic. It includes different clades, some of which have been recently proposed as new genera ( Sharp et al. 2009, Engene et al. 2010, 2013, Komárek et al. 2013, McGregor and Sendall 2014, Stoyanov et al. 2014, Caires et al. 2018). According to Engene et al. (2011), the fact that these unrelated lineages possess highly comparable morphological features is, most likely, the result of convergent evolution, perhaps as a consequence of these species occupying similar ecological niches. According to Komárek (2018), many morphological markers, which are traditionally used for the delimitation of taxa, are very polymorphic, because of this the molecular approach must be used as the primary condition for classification of organisms.
Capillus gen. nov. formed a well-supported clade in our phylogenetic analyses, separated from other clades of marine species, such as Okeania, Moorea and Lyngbya confervoides (here named as ‘True Lyngbya’ clade), which represents the type species of Lyngbya, as stated by Komárek et al. (2013). There are no sequences of L. confervoides from Cádiz (Spain), which is indicated as type locality of this type species ( Lipkin and Silva 2002). All sequences of L. confervoides included in GenBank are from Florida (USA). Cádiz and Florida are warm localities of temperate region. According to Gomont (1892) and Komárek and Anagnostidis (2005), L. confervoides presents large distribution, mainly in temperate zones. This distribution together with our results can indicate that L. confervoides occurs only in temperate regions, and similar populations of this species referred to tropical zones, possibly belong to other genera.
Capillus sp. 2.1, which is positioned as a sister to C. salinus sp. nov., is a further new species to be morphologically and ultrastructurally described. It was isolated from a bank of a river with highly saline water in India ( Mukherjee et al. 2015). Although these two species of this subclade occur in halophilic environments, certainly the salinity in the river is lower than that from marine tidal pools where we collected C. salinus. Sequences in the subclade Capillus sp. (accession numbers from GenBank: NR114680, AJ000714, NR112110, and AB039013) are from the same reference strain PCC 7419, which was collected in a salt marsh ( Castenholz et al. 2001). The use of different methods for obtaining these sequences by each author can explain the few divergences among them observed in our phylogenetic analyses.
Our phylogeny has exhibited five clades containing sequences designate as Lyngbya, besides the clade named ‘True Lyngbya’ which includes sequences of L. confervoides. In addition to these clades, four sequences scattered throughout the phylogenetic reconstruction are also tagged as Lyngbya (underline sequences) ( Fig. 4). The simple morphologic pattern presented by Lyngbya species is easily confused with other genera into Oscillatoriaceae, like genus Oscillatoria Vaucher ex Gomont, which presents similar morphologic pattern, or Phormidium Kützing ex Gomont which can have sheath under specific environmental conditions, as mentioned by Komárek (2018). Consequently, the wrong identification and designation of the strains / sequences can lead to different phylogenetic lineages tagged with the same generic name. Another possibility to explain the significative number of Lyngbya sequences spread in our phylogeny is that the genetic diversity into this genus is greater than phenotypic one, as demonstrated by our results. The ITS region links 16S and 23S rRNA, and it is considered less conserved than 16S rRNA. Moreover, its configuration has been used in a number of studies for species delimitation ( Boyer et al. 2001, 2002, Iteman et al. 2002, Johansen et al. 2011, Engene et al. 2012, Mühlsteinová et al. 2014, Vaz et al. 2014, Martins and Branco 2016, Sciuto and Moro 2016, Brito et al. 2017, Caires et al. 2018). Configuration of ITS regions of C. salinus sp. nov. and C. tropicalis exhibited slight differences in length and configuration of the V3 helix, which did not help in their specific delimitation. Box B helices were very different between the new species, corroborating their separation in different taxa, and agreeing with our phylogenetic analyses, in which these species were well delimitated in the Capillus gen. nov. clade. The new species presented some divergences in D1-D1′ bulges and in basal stems, which were also different from the one observed in the related taxon Limnoraphis sp. (JX006094 and JX006095). According to Reháková et al. (2007), Genuário et al. (2015), and Hentschke et al. (2016, 2017), the number of nucleotides opposing the first lateral bulge appears to be conserved among genera and is useful to separate taxa, reinforcing the description of the genus Capillus, as well as the separation of its two species. This study helps to better understand the diversity of Lyngbya-like cyanobacteria and shows how the biodiversity of benthic filamentous cyanobacteria on tropical coastal environments is poorly explored. Brazilian coastal populations of Cyanobacteria remain largely understudied, hence, it is necessary to increase sampling in this region, as well as using the polyphasic approach in the future studies. It is also important to expand the research tools, comprising morphological, ecological, and molecular data in an integrated taxonomic approach. Such approach will contribute to more accurate taxon classification and biodiversity estimation in cyanobacterial classification systems.
ACKNOWLEDGEMENTS
This study, including a scholarship to TAC (141474/2013-0), was supported by Conselho Nacional de Pesquisa (CNPq–UNIVERSAL 477614/2013-2), Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB–PRONEM PNE 0020/2011; T.O. RED 006/2012), and Instituto Gonçalo Muniz (FIOCRUZ-BA). JMCN acknowledges CNPq fellowship (307368/2015-7). We thank Dra. Alessandra Selbach Schnadelbach (UFBA) and Dr. Eduardo Mendes da Silva (UFBA) for the use of their laboratory facilities. We also thank Dra. Marli de Fátima Fiori (CENA-USP/Piracicaba) and Dra. Ana Paula Dini Andreote (CENA-USP/Piracicaba) for helping us with the initial molecular analyses.
Fig. 1
Map of the Brazilian coast showing collection sites. 
Fig. 2
Capillus salinus sp. nov. ALCB 114379. (A–C) Epilithic population in tidal pool. (D) Fasciculate thallus. (E–L) Photomicrographs. (E) General aspect of filaments and necridic cells (arrow). (F) Cross-walls granulated. (G–I) Different shapes of apical cells. (J & K) Diagonal fragmentation of trichome (arrows). (L) Hormogone. (M–P) Scanning electron microscopy images. (M) Filaments. (N) Detail of trichome. (O) Apical region of trichome. (P) Hormogone release. (Q–U) Ultrastructural sections (transmission electron microscopy). (Q) General aspect of trichome and mucilaginous sheath (ms). (R) Irregular thylakoids (th) arrangement. cr, cross-walls. (S) Cyanophycin granules (cy) placed near the cross-walls (cr). (T) Gas vesicles (gv) distributed in inter-thylakoid spaces and near the cell walls (cw). (U) Detail of gas vesicles (gv). Scale bars represent: D, 400 μm; E–N & P, 10 μm; O, 5 μm; Q, 2 μm; R, S & U, 0.2 μm; T, 0.5 μm. [Colour figure can be viewed at http://www.e-algae.org]. 
Fig. 3
Characterization of Capillus tropicalis sp. nov. ALCB 114392. (A) Epilithic populations in protected reef region. (B–G) Photomicrographs. (B) Apical cell. (C) Granulated cross-walls. (D) Diagonal fragmentation of trichome (arrow). (E) Contorted trichome. (F & G) Necridial regions (arrow indicates crosswise fragmentation of trichome). (H) Scanning electron microscopy image. Detail of trichome and sheaths. (I & J) Ultrastructural sections (transmission electron microscopy). (I) General aspect of trichome. (J) Detail of gas vesicles (gv) nearby cross-walls (cr) and irregular arrangement of thylakoids (th). (K) Cyanophycin granules (cy). Scale bars represent: A, 1.5 cm; B–H, 10 μm; I, 1 μm; J, 0.2 μm; K, 0.5 μm. [Colour figure can be viewed at http://www.e-algae.org]. 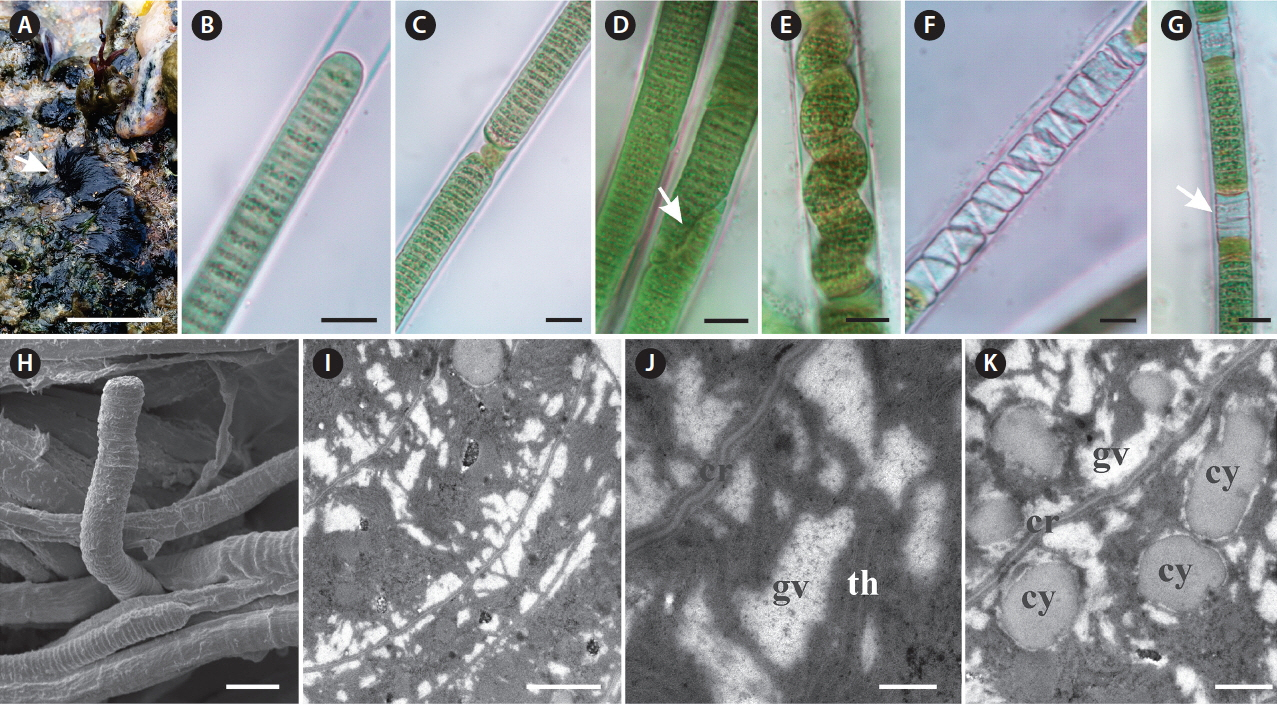
Fig. 4
Maximum likelihood (ML) phylogenetic tree based on the 16S rRNA gene sequences (ntax = 97). Support values >75% are indicated. Values are ML bootstrap (left) and Bayesian posterior probabilities converted to percentages (right). Asterisks indicate that the node is supported at 100%. The new sequences produced in this study are in boldface font; other sequences from GenBank (names are followed by accession numbers). Lyngbya sequences spreaded along the phylogenetic tree are underlined. Bar represents: 0.04 substitutions per nucleotide position. OTUs, operational taxonomic unit. [Colour figure can be viewed at http://www.e-algae.org]. 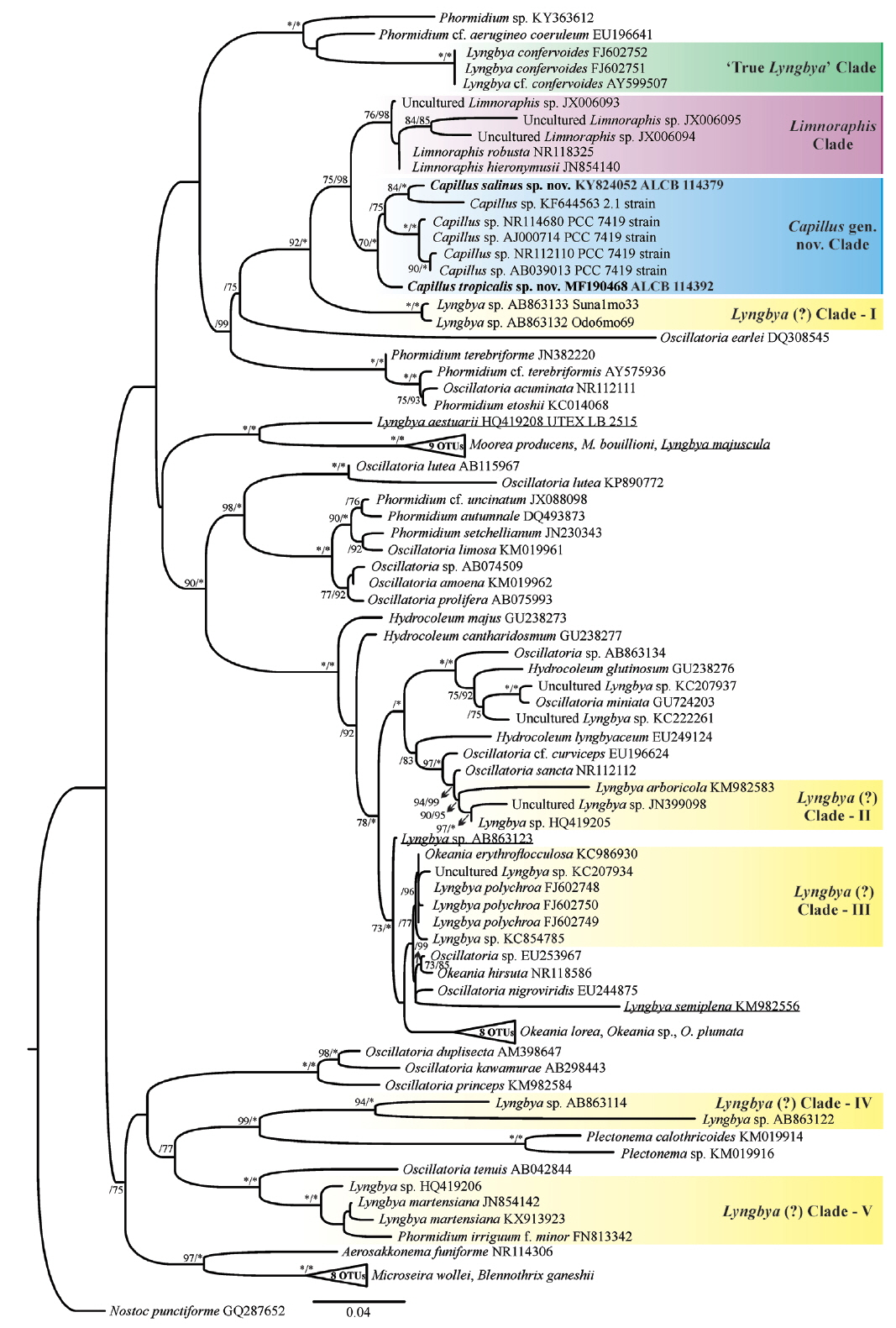
Fig. 5
Secondary structures of 16S–23S internal transcribed spacer sequences of Capillus salinus sp. nov., C. tropicalis sp. nov., and related taxon Limnoraphis sp. (JX006094 and JX006095). [Colour figure can be viewed at http://www.e-algae.org]. 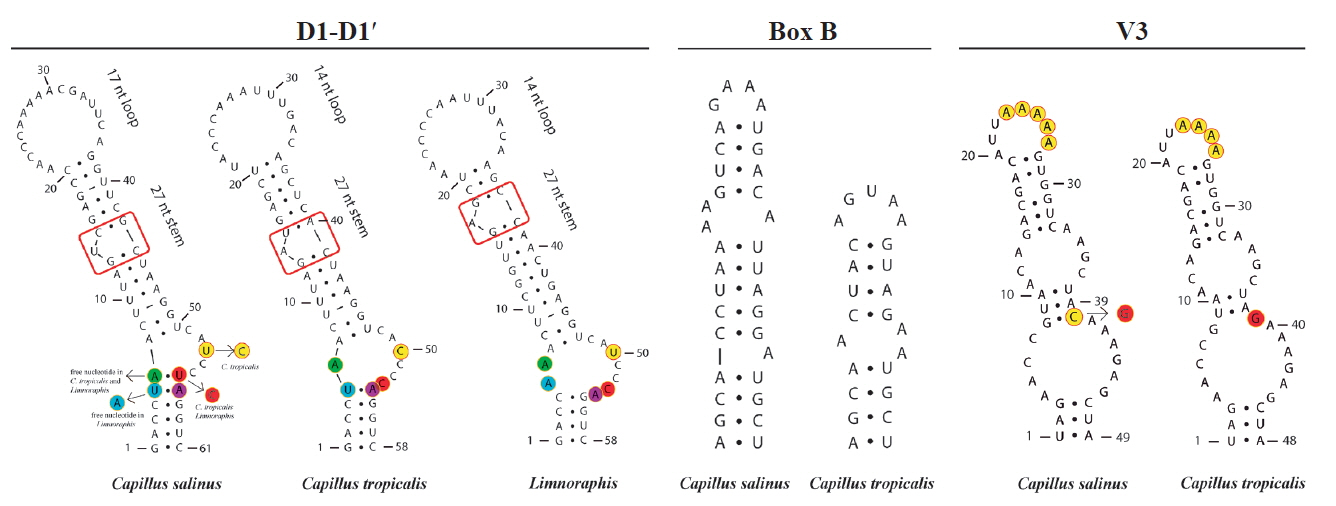
Table 1
Comparison among Capillus gen. nov. and morphologically similar genera
|
Characteristic |
Limnoraphis
|
Lyngbya
|
Capillus gen. nov. |
|
Thallus |
Filaments solitary, free-floating, or growing in small aggregations |
Mainly forming thin or thick compact, large, layered, leathery prostate mats, rarely solitary |
Thallus forming extensive fasciculate mats or small clusters |
|
Filament |
5–25 μm diam.; straight or slightly curved |
Mainly wider than 6 μm diam.; straight or slightly waved |
(8-) 12–30 (−46) μm diam.; straight |
|
Sheath |
Firm, colorless, hyaline, thin, or slightly thick |
Occurs ±obligatorily, firm, thin or thick, colorless, or slightly yellow-brown or reddish, sometimes slightly lamellated |
Hyaline or bright yellow to yellow-brown, firm, thin or thick, sometimes lamellated |
|
Trichome width |
Not available |
Not available |
(6.6-) 10–21 (−24) μm |
|
Cell content |
Gas vesicles facultative |
Without aerotopes, or rarely with gas vesicles in planktic species |
Containing numerous gas vesicles usually forming aerotopes |
|
Constriction |
No or slightly constricted |
Not constricted or constricted |
Not or slightly constricted |
|
Cross-wall |
Not available |
Not available |
Often granulated |
|
Attenuation |
Not attenuated |
Not available |
Not or slightly attenuated towards ends |
|
Cells |
Short, always larger than long |
Short, up to 15 times larger than long |
Short, 1.4–5.6 (−6) μm long, 1.2–11 times larger than long |
|
Apical cell |
Without clear calyptra |
Usually with thickened or with calyptra |
Rounded, conical, conical-rounded, flat-rounded, truncate, rarely capitate, sometimes with thickened, without calyptra |
|
Reproduction |
Intense hormogonia production |
By trichome disintegration into ±short motile hormogonia by the help of necridia; straight fragmentation’s pattern |
Hormogonia formed by straight or diagonal fragmentation of the trichome, helped or not by necridic cells |
|
Occurrence |
Mesotrophic waters in the Lake Atitlán, Guatemala |
Distributed in marine, halophilic, freshwater, and subaerophytic habitats |
Marine and halophilic, and in inland saline biotopes |
|
Reference |
Komárek et al. (2013)
|
Komárek and Anagnostidis (2005)
|
This study |
REFERENCES
Baeta-Neves, MHC. 1991. Estudo das Cianofíceas Marinhas Bentônicas da Região de Cabo Frio (Estado do Rio de Janeiro, Brasil). II–Hormogonae Rev Bras Biol. 52:641–659.
Boyer, SL., Johansen, JR., Flechtner, VR. & Howard, GL. 2002. Phylogeny and genetic variance in terrestrial Microcoleus (Cyanophyceae) species based on sequence analysis of the 16S rRNA gene and associated 16S–23S ITS region. J Phycol. 38:1222–1235.  Brito, A., Ramos, V., Mota, R., Lima, S., Santos, A., Vieira, J., Vieira, CP., Kaštovský, J., Vasconcelos, VM. & Tamagnini, P. 2017. Description of new genera and species of marine cyanobacteria from the Portuguese Atlantic coast. Mol Phylogenet Evol. 111:18–34.   Caires, TA., de Mattos Lyra, G., Hentschke, GS., de Gusmão Pedrini, A., Sant’Anna, CL. & de Castro Nunes, JM. 2018.
Neolyngbya gen. nov. (Cyanobacteria, Oscillatoriaceae): a new filamentous benthic marine taxon widely distributed along the Brazilian Coast. Mol Phylogenet Evol. 120:196–211.   Caires, TA., Sant’Anna, CL. & de Castro Nunes, JMC. 2013. A new species of marine benthic cyanobacteria from the infralittoral of Brazil: Symploca infralitoralis sp. nov. Braz. J Bot. 36:159–163.
Castenholz, RW., Rippka, R. & Herdman, M. 2001. Form-genus VII. Lyngbya Oxygenic photosynthetic bacteria. In : Boone DR, Castenholz RW, Garrity GM, editors Bergey’s Manual of Systematic Bacteriology. Springer, New York, 473–599.
Crispino, LMB. & Sant’Anna, CL. 2006. Cianobactérias marinhas bentônicas de ilhas costeiras do Estado de São Paulo, Brasil. Rev Bras Bot. 29:639–656.   Drummond, AJ., Ashton, B., Buxton, S., Cheung, M., Cooper, A., Duran, C., Field, M., Heled, J., Kearse, M., Markowitz, S., Moir, R., Stones-Havas, S., Sturrock, S., Thierer, T. & Wilson, A. 2011. Geneious v6.0.6, Available from: http://www.geneious.com
. Accessed Oct 8, 2017 Engene, N., Choi, H., Esquenazi, E., Rottacker, EC., Ellisman, MH., Dorrestein, PC. & Gerwick, WH. 2011. Underestimated biodiversity as a major explanation for the perceived rich secondary metabolite capacity of the cyanobacterial genus Lyngbya
. Environ Microbiol. 13:1601–1610.    Engene, N., Coates, RC. & Gerwick, WH. 2010. 16S rRNA gene heterogeneity in the filamentous marine cyanobacterial genus Lyngbya
. J Phycol. 46:591–601.  Engene, N., Rottacker, EC., Kaštovský, J., Byrum, T., Choi, H., Ellisman, MH., Komárek, J. & Gerwick, WH. 2012.
Moorea producens gen. nov., sp. nov. and Moorea bouillonii comb. nov., tropical marine cyanobacteria rich in bioactive secondary metabolites. Int J Syst Evol Microbiol. 62(Pt 5):1171–1178.    Fiore, MF., Moon, DH., Tsai, SM., Lee, H. & Trevors, JT. 2000. Miniprep DNA isolation from unicellular and filamentous cyanobacteria. J Microbiol Methods. 39:159–169.   Genuário, DB., Vaz, MGMV., Hentschke, GS., Sant’Anna, CL. & Fiore, MF. 2015.
Halotia gen. nov., a phylogenetically and physiologically coherent cyanobacterial genus isolated from marine coastal environments. Intern J Syst Evol Microbiol. 65:663–675.  Gomont, M. 1892. Monographie des Oscillariées (Nostocacées Homocystées). Deuxième partie. - Lyngbyées
. Ann Sci Nat Bot Sér 7. 16:91–264.
Hentschke, GS., Johansen, JR., Pietrasiak, N., Fiore, MF., Rigonato, J., Sant’Anna, CL. & Komárek, J. 2016. Phylogenetic placement of Dapisostemon gen. nov. and Streptostemon, two tropical heterocytous genera (Cyanobacteria). Phytotaxa. 245:129–143.  Hentschke, GS., Johansen, JR., Pietrasiak, N., Rigonato, J., Fiore, MF. & Sant’Anna, CL. 2017.
Komarekiella atlantica gen. et sp. nov. (Nostocaceae, Cyanobacteria): a new subaerial taxon from the Atlantic Rainforest and Kauai, Hawaii. Fottea. 17:178–190.   Iteman, I., Rippka, R., Tandeau de Marsac, N. & Herdman, M. 2002. rDNA analyses of planktonic heterocystous cyanobacteria, including members of the genera Anabaenopsis and Cyanospira
. Microbiology. 148(Pt 2):481–496.   Jacinavicius, FR., Gama, WA. Jr, Azevedo, MTP. & Sant’Anna, CL. 2012. Manual para cultivo de cianobactérias. Publicações Online do Instituto de Botânica de São Paulo, São Paulo, 32 pp.
Johansen, JR., Kovacik, L., Casamatta, DA., Fučiková, K. & Kaštovský, J. 2011. Utility of 16S–23S ITS sequence and secondary structure for recognition of intrageneric and intergeneric limits within cyanobacterial taxa: Leptolyngbya corticola sp. nov. (Pseudanabaenaceae, Cyanobacteria). Nova Hedwigia. 92:283–302.  Karnovsky, MJ. 1965. A formaldehyde-glutaraldehyde fixative of high osmolarity for use in electron microscopy. J Cell Biol. 27:1A–149A.  Komárek, J. 2016. A polyphasic approach for the taxonomy of cyanobacteria: principles and applications. Eur J Phycol. 51:346–353.  Komárek, J. 2018. Delimitation of the family Oscillatoriaceae (Cyanobacteria) according to the modern polyphasic approach (introductory review). Braz J Bot. 41:449–456.   Komárek, J. & Anagnostidis, K. 2005. Cyanoprokaryota-2 Teil/2nd part: Oscillatoriales. In : Büdel B, Gärtner G, Krienitz L, Schagerl M, editors Süsswasserflora von Mitteleuropa 19⁄778 2. Elsevier/Spektrum, Heidelberg, 1–759.
Komárek, J., Kaštovský, J., Mareš, J. & Johansen, JR. 2014. Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) 2014, using a polyphasic approach. Preslia. 86:295–335.
Komárek, J., Zapomělová, E., Šmarda, J., Kopecký, J., Rejmánková, E., Woodhouse, J., Neilan, BA. & Komárková, J. 2013. Polyphasic evaluation of Limnoraphis robusta, a water-bloom forming cyanobacterium from Lake Atitlán, Guatemala, with a description of Limnoraphis gen. nov. Fottea. 13:39–52.   Lane, DJ. 1991. 16S/23S rRNA sequencing. In : Stackebrandt E, Goodfellow M, editors Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons, Chichester, 115–175.
Lipkin, Y. & Silva, PC. 2002. Marine algae and seagrasses of the Dahlak Archipelago, southern Red Sea. Nova Hedwigia. 75:1–90.  Martins, MD. & Branco, LHZ. 2016.
Potamolinea gen. nov. (Oscillatoriales, Cyanobacteria): a phylogenetically and ecologically coherent cyanobacterial genus. Int J Syst Evol Microbiol. 66:3632–3641.   Mau, B., Newton, MA. & Larget, B. 1999. Bayesian phylogenetic inference via Markov chain Monte Carlo methods. Biometrics. 55:1–12.   McGregor, GB. & Sendall, BC. 2014. Phylogeny and toxicology of Lyngbya wollei (Cyanobacteria, Oscillatoriales) from north-eastern Australia, with a description of Microseira gen. nov. J Phycol. 51:109–119.   Mühlsteinová, R., Johansen, JR., Pietrasiak, N., Martin, MP., Osorio-Santos, K. & Warren, SD. 2014. Polyphasic characterization of Trichocoleus desertorum sp. nov. (Pseudanabaenales, Cyanobacteria) from desert soils and phylogenetic placement of the genus Trichocoleus
. Phytotaxa. 163:241–261.  Mukherjee, C., Chowdhury, R., Sutradhar, T., Begam, M., Ghosh, SM., Basak, SK. & Ray, K. 2015. Parboiled rice effluent: a wastewater niche for microalgae and cyanobacteria with growth coupled to comprehensive remediation and phosphorus biofertilization. Algal Res. 19:225–236.  Neilan, BA., Jacobs, D., Del Dot, T., Blackall, LL., Hawkins, PR., Cox, PT. & Goodman, AE. 1997. rRNA sequences and evolutionary relationships among toxic and nontoxic cyanobacteria of the genus Microcystis
. Int J Syst Bacteriol. 47:693–697.   Nylander, JAA. 2008. MrModeltest 23 Program distributed by the author. Evolutionary Biology Centre. Uppsala University.
Porta, D. & Hernández-Mariné, M. 2005. Structural and ultrastructural characterization of several culture strains assigned to Oscillatoria and Lyngbya (Cyanophyta/Cyanoprokaryota/Cyanobacteria). Algol Stud. 117:349–370.  Reháková, K., Johansen, JR., Casamatta, DA., Xuesong, L. & Vincent, J. 2007. Morphological and molecular characterization of selected desert soil cyanobacteria: three species new to science including Mojavia pulchra gen. et sp. nov. Phycologia. 46:481–502.  Rippka, R., Deruelles, J., Waterbury, JB., Herdman, M. & Stanier, RY. 1979. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol. 111:1–61.  Ronquist, F., Teslenko, M., van der Mark, P., Ayres, DL., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, MA. & Huelsenbeck, J. 2012. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 61:539–542.     Sambrook, JF. & Russell, DW. 2001. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York, 2100 pp.
Sant’Anna, CL. 1995. Cyanophyceae marinhas bentônicas do Parque Estadual da Ilha do Cardoso, SP, Brasil. Hoehnea. 22:197–216.
Sant’Anna, CL. 1997. Cyanophyceae marinhas bentônicas da região de Ubatuba, SP, Brasil. Hoehnea. 24:57–74.
Sant’Anna, CL., Cordeiro-Marino, M., Braga, MRA. & Guimarães, SMPB. 1985. Cianofíceas marinhas bentônicas das Praias de Peruíbe e dos Sonhos, Município de Itanhaém, São Paulo, Brasil. Rickia. 12:89–112.
Sciuto, K. & Moro, I. 2016. Detection of the new cosmopolitan genus Thermoleptolyngbya (Cyanobacteria, Leptolyngbyaceae) using the 16S rRNA gene and 16S–23S ITS region. Mol Phylogenet Evol. 105:15–35.   Sharp, K., Arthur, KE., Gu, L., Ross, C., Harrison, G., Gunasekera, SP., Meickle, T., Matthew, S., Luesch, H., Thacker, RW., Sherman, DH. & Paul, VJ. 2009. Phylogenetic and chemical diversity of three chemotypes of bloom-forming Lyngbya species (Cyanobacteria: Oscillatoriales) from reefs of southeastern Florida. Appl Environ Microbiol. 75:2879–2888.    Vaz, MGMV., Genuário, DB., Andreote, APD., Malone, CFS., Sant’Anna, CL., Barbiero, L. & Fiore, MF. 2014.
Pantanalinema gen. nov. and Alkalinema gen. nov.: novel pseudanabaenacean genera (Cyanobacteria) isolated from saline–alkaline lakes. Int J Syst Evol Microbiol. 65:298–308.   Werner, VR., Cabezudo, MM., Neuhaus, EB., Caires, TA., Sant’Anna, CL., Azevedo, MTP., Malone, C., Gama, WA. Jr, Santos, KRS. & Menezes, M. 2017. Cyanophyceae in Lista de Espécies da Flora do Brasil. Jardim Botânico do Rio de Janeiro, Available from: http://floradobrasil.jbrj.gov.br/jabot/floradobrasil/FB108025
. Accessed Aug 27, 2017
|
|
















