INTRODUCTION
Red rot disease was first reported on Pyropia tenera in Japan (Arasaki 1947), and its pathogenesis has been characterized in Pyropia species (Takahashi et al. 1977). Outbreaks of red rot disease on Pyropia species have caused major damage to commercial Pyropia aquaculture systems in Korea, Japan, and China (Kawamura et al. 2005, Blouin et al. 2011, Kim et al. 2014).
After the first report of red rot disease on Pyropia tenera, only Pythium porphyrae (Pythiales, Oomycetes) was identified as a pathogen of red rot disease in Pyropia species in Korea, Japan, and China (Park et al. 2006, Liu et al. 2012, Kim et al. 2014). Recently, we isolated Pythium chondricola from the infected blades of Pyropia yezoensis collected from commercial Pyropia aquaculture farms based on the mitochondrial cox1 region (Lee et al. 2015).
Using cox1 markers, P. chondricola was detected in Py. yezoensis, and was confirmed to be genetically different from the P. porphyrae detected in Japan. This was the first report of the presence of P. chondricola on Pyropia species, following the first taxonomic report of its occurrence on the red alga Chondrus crispus (De Cock 1986). However, the pathogenesis of P. chondricola must be proven to confirm that it is an algal pathogen causing red rot disease in Pyropia species (i.e., proven to satisfy Koch’s postulates).
For precise identification of oomycete species, it is necessary to develop multiple molecular markers (Lévesque and De Cock 2004, Seifert 2009, Robideau et al. 2011). In this regard, Choi et al. (2015) recently compared the efficiency of using the pattern of sequence variation between cox2 and cox1 regions for species identification. They accordingly found that cox2 showed higher interspecific and lower intraspecific divergence than cox1, and was more successful in species identification. Therefore, the cox2 region has been suggested as a successful partner DNA barcode marker.
Following the detection of P. chondricola in an infected blade of Py. yezoensis using the cox1 region (Lee et al. 2015), a new cox2 gene sequence (KJ595354) of P. chondricola was published in GenBank (National Center for Biotechnology Information, NCBI). Therefore, we developed a new primer set to amplify the cox2 region that also exhibits specificity for Pythium species. A molecular examination was conducted based on the Pythium-specific cox2 gene marker developed in this study. Pythium cox2 sequences were isolated from the infected blades of Py. yezoensis with red rot disease, and were compared with cox2 sequences in GenBank. We also conducted a re-inoculation test to satisfy Koch’s postulates using an isolate from the infected Py. yezoensis. These analyses were expected to confirm the pathogenesis of P. chondricola on Py. yezoensis.
MATERIALS AND METHODS
Sample preparation
We used Pythium-infected blades of Py. yezoensis collected from aquaculture farms in Korea (Biando [Gunsan] and Aphaedo [Shinan] in December 2014). Pyropia samples with signs of red rot disease were also collected from Gaeyado (Dec 2014, Gunsan, Korea). The presence of P. chondricola was confirmed in these Pyropia samples (Lee et al. 2015). Strains of P. chondricola were isolated from Py. yezoensis collected from Biando and Aphaedo (Dec 2014, Korea) and were cultured in the Seaweed Research Center (National Institute of Fisheries Science, Mokpo, Korea). In addition, a culture strain of P. chondricola (Biando, Korea; NIFS-PC-001) has been deposited in the Marine BioResources Bank (MBRB) in the Seoul National University, Korea (SFC20170403-M01). Species identification of the cultured strain was conducted using the cox1 marker (Lee et al. 2015).
Re-inoculation experiment
We inoculated the P. chondricola strain onto cornmeal agar and Arasaki B medium (Arasaki et al. 1968), and maintained the culture (Biando) at 15°C. A re-inoculation test was performed in accordance with Koch’s postulates. Blades of Py. yezoensis used in the re-inoculation test were cultured at 15°C under white fluorescent irradiation of approximately 20 μmol photons m−2 s−1 and a 12 : 12-h light : dark cycle. Morphological observations during the infection of P. chondricola on the healthy blade of Py. yezoensis were conducted using a microscope (Nikon Eclipse Ni-u, Tokyo, Japan) (Fig. 1).
Development of a Pythium-specific cox2 marker and molecular analyses
We developed a new primer set to amplify the cox2 region of P. chondricola. For primer design, we compared two cox2 gene sequences of P. chondricola (KJ595354) and P. porphyrae (KJ595377) from GenBank. We initially searched the conserved regions of the two species. We excluded the cox2 region conserved among other oomycetes, except for Pythium species.
Because these two species have a close taxonomic relationship (Lévesque and De Cock 2004), a new cox2 marker was designed for putative specificity in P. chondricola, P. porphyrae, and P. adhaerens. To evaluate the specificity of the newly developed cox2 primers for Pythium species, we compared their specificity against that of previously reported cox2 primer pairs (COX2F/COX2R, FM66/FM58) (Hudspeth et al. 2000, Villa et al. 2006, Senda et al. 2009, Choi et al. 2015).
DNA extraction, polymerase chain reaction (PCR) amplification, and sequencing processes followed the methods described by Lee et al. (2015). Total DNAs were extracted from a culture strain of P. chondricola and the infected field samples of Py. yezoensis. The putative infection of P. chondricola in Pyropia samples was investigated using the cox1 marker (Lee et al. 2015). Similarity analysis was conducted using the BLAST searching tool in GenBank.
RESULTS
Infection of Pyropia yezoensis by Pythium chondricola
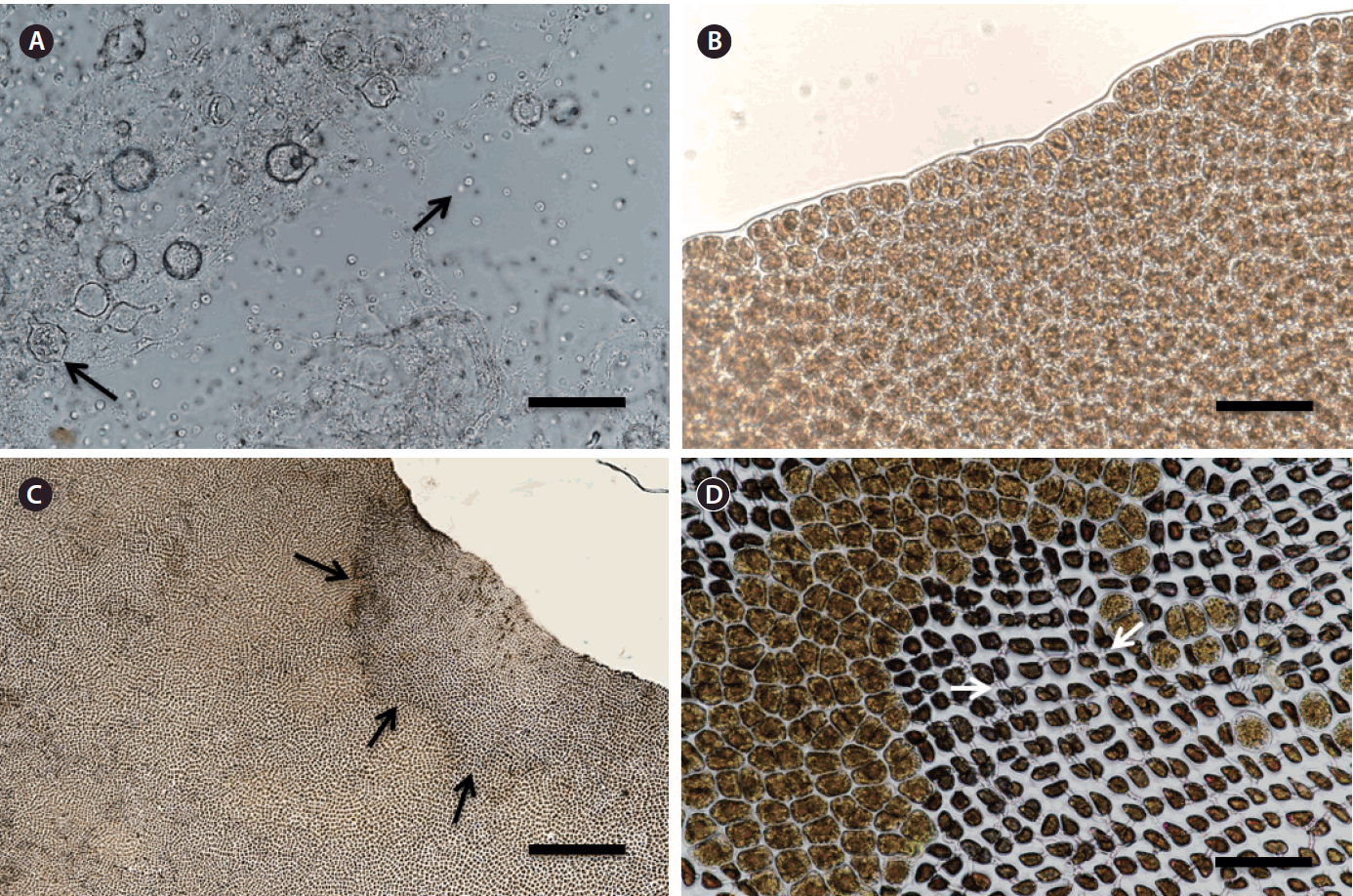
The morphological features of Py. yezoensis blades were observed during infection by P. chondricola. The formation of a zoosporangium and the release of zoospores were accordingly observed (Fig. 1A). Two days after re-inoculation, Pyropia cells in the blade showed symptoms of red rot disease. The area of the lesion in the infected blade lost its original color and developed a purple and greenish color (Fig. 1C). P. chondricola mycelia were also observed in the lesioned area (Fig. 1D).
Pythium-specific cox2 marker development for the detection of Pythium chondricola
In this study, we developed the following primer pair: forward primer, cox2-PACP-F1 (5′-GATGTTATTTAAAACAATAGTTC-3′); and reverse primer, cox2-PACP-R1 (5′-TAAAGAAGGAATAGCCCAA-3′). PCR was successfully conducted on the infected blades of Py. yezoensis using this new primer pair. We also acquired PCR products from the culture strain. The COX2F/COX2R and FM66/FM58 primer pairs used in previous studies showed amplicons of different sizes, approximately 600 bp.
From sequencing, we successfully isolated the cox2 region using our newly developed primer pair (cox2-PACP-F1/cox2-PACP-R1). The cox2 sequences isolated from culture strains of P. chondricola and the infected host blades of Py. yezoensis (Biando and Aphaedo) have an identical sequence (GenBank accession number KU754146 [373 bp]). With the exception of the primer binding sites, the cox2 regions showed 100% similarity (373/373) with that P. chondricola (KJ595354; CBS:203.85, The Netherlands), 98% (367/373) with that of P. porphyrae (KJ595377; CBS:369.79, Japan), and 97% (362/373) with that of P. adhaerens (KJ595386; CBS:520.74, The Netherlands). The cox1 sequences of these three Pythium strains (CBS:203.85, CBS:369.79, CBS:520.74) were also found in GenBank, and these sequences were analyzed by Lee et al. (2015). We have deposited the cox1 sequences of P. chondricola reported by Lee et al. (2015). All Korean samples have an identical cox1 sequence (GenBank accession number KY124355, 386 bp). Whereas the interspecific sequence variations identified using the cox1 region were consistent, the cox2 marker showed more variable sequences.
The primers used in previous cox2 studies could not amplify the cox2 region of Pythium species. The primer pairs COX2F/COX2R (631 bp) and FM66/FM58 (613 bp) amplified the cox2 region of the host, Py. yezoensis. The sequence of the amplicons had 100% similarity with the cox2 sequence of Py. yezoensis (JQ736809; each 581 bp and 573 bp).
DISCUSSION
The re-inoculation test successfully confirmed that the isolated P. chondricola causes pathogenesis in Py. yezoensis. The infection by P. chondricola caused cell death in healthy blades of Py. yezoensis maintained under indoor culture. The infected blades of Py. yezoensis showed characteristics typical of red rot disease (Takahashi et al. 1977).
We previously isolated cox1 and internal transcribed spacer (ITS) sequences from culture strains of P. chondricola and infected samples of Py. yezoensis collected from aquaculture farms (Lee et al. 2015). However, the ITS sequences failed to discriminate P. chondricola from P. porphyrae. In contrast, the cox1 region successfully provided useful genetic information for species identification. It has been suggested that a recently identified oomycete DNA barcode (Choi et al. 2015) could be used with the cox2 region, serving as a partner DNA barcode, because of its efficiency for PCR and sufficient variation for species identification. The cox2 region from the culture strain of P. chondricola is identical to that from the infected blade of Py. yezoensis.
The cox2 region has been used for molecular taxonomic studies of oomycetes (Hudspeth et al. 2000, Villa et al. 2006, Senda et al. 2009), and effective primer combinations were also developed. In these studies, most of the DNA samples were extracted from Pythium culture strains, and not from environmental samples such as infected Pyropia blades from aquaculture farms. Therefore, in the case of DNA analysis for host species of pathogens, the primer pair used to amplify the cox2 region should be verified to specify the target pathogen species.
In this study, we assessed the efficacy of universal primers of the cox2 region for oomycetes to isolate the cox2 region from the infected blades of Py. yezoensis. However, we were unable to acquire the Pythium cox2 sequence. Furthermore, the universal primers did not show specificity for Pythium species, since this primer pair also amplified the cox2 region of the host Py. yezoensis. Therefore, cox2 primers that are appropriate for broad taxonomic groups of oomycetes might fail to detect the presence of Pythium species in the infected blades of Py. yezoensis.
We developed a new cox2 primer set that exhibited specificity for Pythium species, thereby enabling us to isolate the Pythium cox2 sequence from host samples showing red rot disease. The primer pair (cox2-PACP-F1/cox2-PACP-R1) developed in this study could successfully amplify the Pythium cox2 sequence with high efficiency and specificity.
For the Py. yezoensis blades collected from Biando and Gaeyado (Gunsan), a universal primer pair showed the non-specific band of Py. yezoensis. Nevertheless, the cox2 primer pair developed in the present study successfully amplified the cox2 region of Pythium from the Biando sample. The sequencing results show that the cox2 amplicons obtained using the universal primer pair exhibited the Py. yezoensis cox2 sequence instead of a Pythium sequence. These results strongly suggest the specificity of this new cox2 primer pair.
To confirm P. chondricola as an algal pathogen, a re-inoculation experiment performed in accordance with Koch’s postulates is required. In addition, the DNA barcode method is needed to detect Pythium species from samples in various states (e.g., decaying host samples or environmental samples, such as seawater and sediment). Here, we examined the pathogenesis of P. chondricola through the re-inoculation test and a novel molecular marker with high efficiency and specificity for Pythium.
Several cox1 sequences have been reported under the names of P. chondricola and P. porphyrae from Japan, Korea, The Netherlands, New Zealand, and the USA (Robideau et al. 2011, Lee et al. 2015, Diehl et al. 2017, Klochkova et al. 2017). The cox1 sequences (HQ708542, HQ708543, KY124355) were identical to that of the type culture of P. chondricola (HQ708544, CBS 203.85). Moreover, three cox1 sequences showed high similarity: HQ708545 (1 bp difference, 679/680 [99%]), KX527563 (2 bp, 576/578 [99%]), and KY650705 (2 bp, 676/678 [99%]). In contrast, the cox1 sequence of the type culture of P. porphyrae (HQ708794; CBS369.79) showed a 7 bp difference (673/680, 99%) from that of the type culture of P. chondricola (HQ708544).
The reported Pythium species show a diverse range of host infection (P. chondricola from Chondrus crispus [red algae], Ulva lactuca [green algae], Zostera marina [flowering plants] in De Cock (1986)); as P. porphyrae (KX527563) on the embryonic roots of various higher plants (e.g., carrot, cucumber, lettuce, and rice) in the artificial infection test of Klochkova et al. (2017); as P. porphyrae (KY650705) on Porphyra and Pyropia species in Diehl et al. (2017); as P. porphyrae on Py. tenera/Py. yezoensis in Kim et al. (2014); and P. chondricola (KY124355) on Py. yezoensis in Lee et al. (2015, this study). Interestingly, this broad range of host infection was also reported from the original description of P. chondricola (De Cock 1986).
Diehl et al. (2017) recently treated P. chondricola as a heterotypic synonym of P. porphyrae based on the identical ITS region and the high sequence similarity of the cox1 region in the two species. However, the close genetic similarity among Pythium species has previously been reported from molecular taxonomic and DNA barcoding studies on the genus Pythium (Lévesque and De Cock 2004, Robideau et al. 2011). On the basis of the phylogenetic analyses of intra/interspecific variations, Robideau et al. (2011) treated P. chondricola as an independent species separate from P. porphyrae. Moreover, they proposed that the cox1 region could be used as a molecular marker to discriminate between P. chondricola and P. porphyrae.
Of course, taxonomic re-examination will need to be conducted to clarify the taxonomic relationship among three allied Pythium species (P. adhaerens, P. chondricola, and P. porphyrae). However, for that purpose, more samples, including type cultures and more DNA sequences, should be examined. Therefore, for further studies, the distinctive genetic features of Korean P. chondricola should be conserved as an independent taxonomic entity apart from P. porphyrae.
The recent findings regarding a novel algal pathogen of Pyropia species causing red rot disease could suggest the cryptic biodiversity of oomycetes pathogens (Mo et al. 2016). Moreover, Park et al. (2003) investigated the genetic variation among Korean and Japanese isolates of P. porphyrae using random amplified polymorphic DNA and detected interesting genetic heterogeneity among Korean isolates. These results also support the possibility of intra/interspecific genetic diversity among Korean Pythium species and the existence of P. chondricola as an independent species. Therefore, this research is expected to provide an accurate and reliable method for monitoring the distribution and infection pattern of P. chondricola/P. porphyrae in Pyropia aquaculture farms.