Verifying a new distribution of the genus Amalthea (Halymeniales, Rhodophyta) with description of A. rubida sp. nov. from Korea
Article information
Abstract
The red algal genus Amalthea was first described from New Zealand with one species, A. freemaniae. We discovered a new species of Amalthea from Korea, Amalthea rubida sp. nov., and described it based on morphology and rbcL sequences. A. rubida is characterized by a foliose, membranous and soft thallus having loosely arranged anticlinal medullary filaments, much like A. freemaniae, except for a difference of the cortex thickness in the basal part of vegetative structure. In the rbcL phylogenetic analyses, A. rubida formed a clade with A. freemaniae from New Zealand and Halymenia abyssicola from Mexico. A. rubida showed 5.7% interspecific divergence to A. freemaniae and 4.7% to H. abyssicola. Our discovery of A. rubida expands the distribution of Amalthea from the oceanic southwestern Pacific into the continental northwestern. A targeted global study is needed to fully reveal the species diversity and phylogenetic relationships of the genus Amalthea and members of the order Halymeniales.
INTRODUCTION
Coasts in different climatic zones, and those situated in climatically similar zones but in different oceans or hemispheres, are mostly inhabited by dissimilar macroalgal seaweed floras (Van den Hoek 1984). In this biogeographic boundaries, the warm and cold temperate regions of the southwestern Pacific have a similarity to the thermogeographic model (Adey and Steneck 2001). The temperate southwestern Pacific regions of Australia and New Zealand have high endemic flora (Phillips 2001, Hurd et al. 2004). However, some global targeted studies has eroded the endemism of southwestern Pacific regions. For example, the genus Psaromenia, thought to be endemic to New Zealand, was found in Korean water (Schneider et al. 2014), indicating that similarities do exist between the flora of the hemispheres.
In foliose red algae, it is difficult to determine their taxonomic position based solely on morphology, due to few taxonomic characters and phenotypic variation. Molecular data is now essential to identify these uncertain entities and has led to the recognition of several foliose red algal genera (D’Archino et al. 2010, 2012, Clarkston and Saunders 2012). Within the order Halymeniales, there are many new genera proposed mostly using molecular data such as Amalthea and Galene (D’Archino et al. 2014), Felicinia (Manghisi et al. 2014), Neorubra (Calderon et al. 2014), and Mariaramirezia (Calderon et al. 2016). Most of these genera still remain monospecific due to the limited sampling or the lack of morphological characters to determine the taxonomic entity.
The genus Amalthea D’Archino et W. A. Nelson was established on the basis of Amalthea freemaniae from New Zealand having foliose, membranous, and soft thallus. A. freemaniae has markedly different vegetative structure in its basal and distal parts. In New Zealand, A. freemaniae was identified as Halymenia latifolia P. Crouan & H. Crouan ex Kützing because of its morphological similarity to Halymenia, i.e., having transverse medullary filaments with stellate or ganglionic cells (D’Archino et al. 2014). The distribution of H. latifolia is mainly localized in Europe with the type locality in Brittany, France (Guiry and Guiry 2016), except for the New Zealand record (Adams 1994). Using phylogenetic analysis of the rbcL gene, D’Archino et al. (2014) showed that A. freemaniae was not related to the genus Halymenia, and sister species to Halymenia abyssicola E. Y. Dawson from the Pacific Mexico (D’Archino et al. 2014).
Investigations of the order Halymeniales in Korea resulted in the recognition of new halymenialean species, including Cryptonemia asiatica M. Y. Yang et M. S. Kim (Yang and Kim 2014), Galene rotunda (Okamura) D’Archino et Zuccarello (as Cryptonemia rotunda, Kim et al. 2012), and Pachymeniopsis volvita M. Y. Yang et M. S. Kim (Yang and Kim 2015). New foliose species remain unrecognized due to their subtidal habitat or similar exterior morphology (Kim et al. 2012). There are 8 genera and 39 species of the order Halymeniales in Korea (Kim et al. 2013). However, foliose species resembling Halymenia, based on similar internal anatomy, have not been reported. The aim of this study is to determine the taxonomic position of foliose specimens with similar anatomy to Halymenia.
MATERIALS AND METHODS
Samples were collected using SCUBA (from depths of 15–20 m) in Dodu and Udo of Jeju Island, Korea from 2015 to 2016. Underwater images and photographs were obtained using a digital camera (G7X; Canon, Tokyo, Japan). Samples for DNA analyses were detached from the thallus and dried in silica-gel. Samples for anatomical investigations were preserved in 5% formalin in seawater. Sections were done by hands using a razor or a bench-top freezing microtome (NK-101-II; Nippon Optical Works Co., Ltd., Tokyo, Japan), stained with 1% acidified aniline blue, and mounted in 30% corn syrup. Photomicrographs were obtained using a BX43 microscope (Olympus, Tokyo, Japan) with an EOS 600D digital camera (Canon). The digitized images were imported into Adobe Photoshop ver. 6.1 (Adobe Systems Inc., San Jose, CA, USA). Voucher specimens were deposited in the herbarium of Jeju National University (JNUB), Korea and the National Institute of Biological Resources (NIBR), Incheon, Korea.
DNA was extracted using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. Following DNA extraction, the rbcL gene was amplified. The primers used to amplify the gene region were F145-R898 and F762-R1442 (Kim et al. 2010). Amplification was performed in a 20 μL reaction using AccuPower PCR PreMix (Bioneer, Daejeon, Korea) and Swift MaxPro thermal cyclers (ESCO, Singapore). The PCR parameters were as follows: initial denaturation at 96°C for 4 min, followed by 35 cycles of denaturation at 94°C for 1 min, annealing at 50°C for 1 min, and extension at 72°C for 2 min with a final extension at 72°C for 7 min. PCR products were purified using the AccuPrep Purification Kit (Bioneer), and sequenced at Macrogen (Seoul, Korea). Sequencing electropherograms were edited using Chromas ver. 1.45 software (Queensland, Australia). Multiple sequence alignments were made in BioEdit software (Hall 1999) and aligned visually.
Uncorrected pairwise distances were estimated using MEGA 4.0 software (Tamura et al. 2011). To construct the rbcL phylogenetic tree, maximum likelihood (ML) analyses were performed using RAxML software (Stamatakis 2006) with the GTR + Γ + I model. To identify the best tree, 200 independent tree inferences using the -# option with default –I (automatically optimized subtree pruning-regrafting rearrangement) and –c (25 distinct rate categories) options were used. To generate bootstrap support values for the phylogeny, 1,000 replicates were implemented. Bayesian phylogenetic inferences (BI) were generated using MrBayes ver. 3.1.2 software (Ronquist and Huelsenbeck 2003). For each matrix, three million generations of two independent runs were performed with four chains and sampling trees every 100 generations. The burn-in period was identified graphically by tracking the likelihood at each generation. The remaining trees were used to calculate a posterior probability tree topology. Aeodes nitidissima J. Agardh (EU497914) and Pachymenia laciniata J. Agardh (AY583301) were used as outgroups.
RESULTS
Eight rbcL sequences were obtained from Korean specimens and aligned with 31 sequences from GenBank. The rbcL data set consisted of 1,518 characters: 431 variable positions (28.4%) and 317 parsimoniously informative positions (20.9%). The phylogenetic trees constructed by ML and BI had the same topology (Fig. 1). Phylogenetic analyses revealed that the Korean specimens formed a clade with the type species A. freemaniae (KJ606651-KJ606653) from New Zealand and H. abyssicola (GU598119) from the Pacific Mexico. The clade was moderately supported, and separated from the other halymenialean genera, including Halymenia, Cryptonemia, and Galene. Eight Amalthea specimens from Korea were identical and had a pairwise distance of 4.7% to H. abyssicola, and 5.7% to A. freemaniae. Our rbcL results show that the samples from Korea are a distinct entity. We described these specimens as a new species of Amalthea.
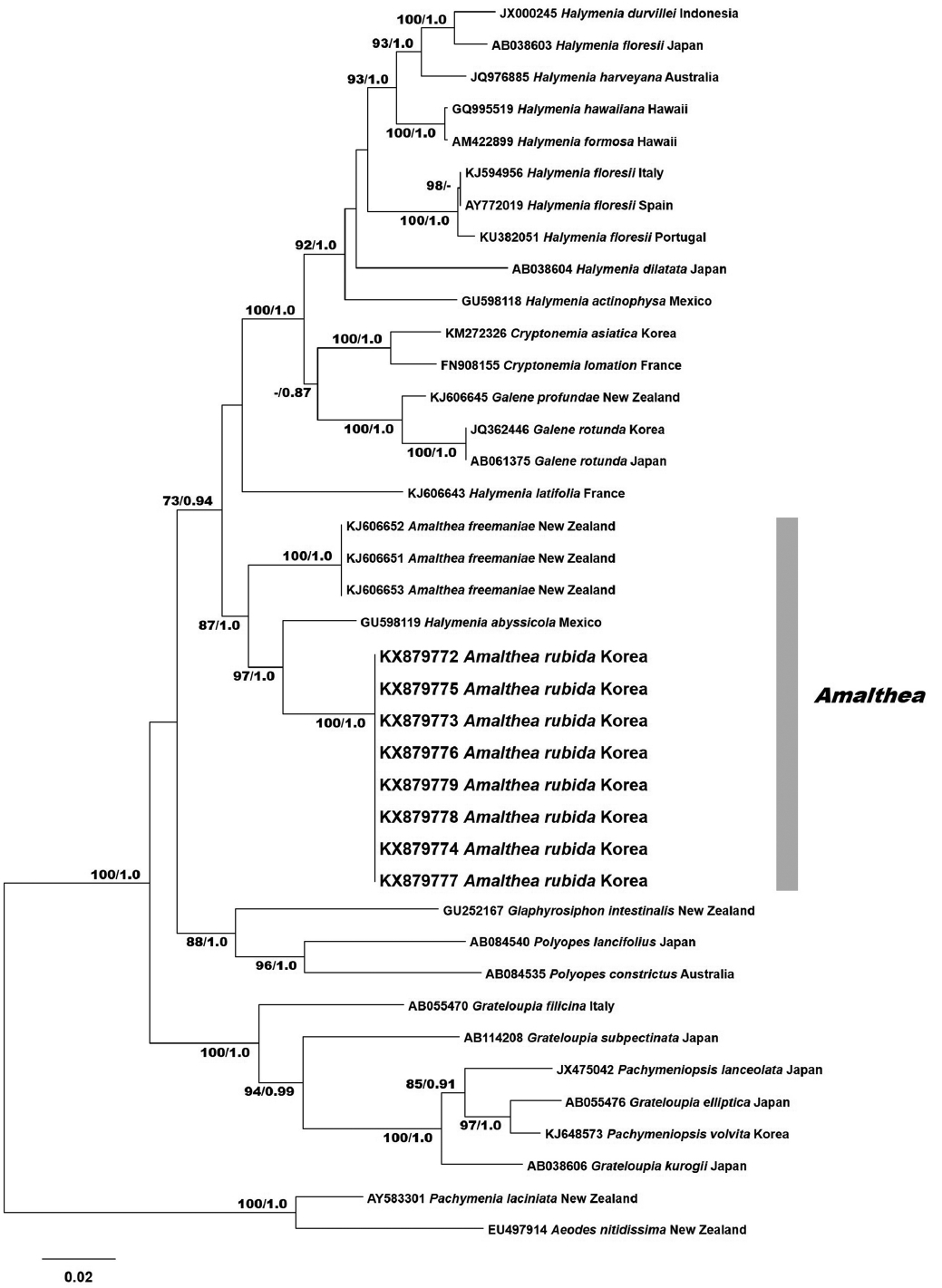
Phylogenetic tree of Amalthea rubida H. W. Lee et M. S. Kim sp. nov. based on rbcL sequences inferred from maximum-likelihood (ML) analysis. Support values at each node are from ML bootstrap (>70%, left) and Bayesian posterior probability (>0.70, right). Scale bar represents: substitution/site.
Amalthea rubida H. W. Lee et M. S. Kim sp. nov
Description
Thallus erect, solitary, foliose, usually entire, occasionally cleft; attached by a small discoid holdfast; growing to 5–15 cm (up to 35 cm) high and 5–10 cm (up to 20 cm) wide (Fig. 2A–C); obovate with cuneate base when young (Fig. 2B & C), expanded irregulary when large (Fig. 2A); epilithic or epiphytic on rhodoliths or crustose coralline algae (Fig. 2D); pale reddish brown to red color (Fig. 2A–D); quite gelatinous, membranous, in the texture; 350–380 μm thick in the basal part and 290–320 μm in the distal part; mostly entire margin, and slightly wavy when large; stipe short, cylindrical to compressed, less than 1 mm long, 480–520 μm in diameter; cortical cells in surface view rounded to sub-rounded, not compact, and 3–5 μm in diameter (Fig. 2E); subcortical stellate cells, 9–12 × 10–15 μm, connected polygonally to each other (Fig. 2F); medullary stellate cells, darkly stained by 1% acidified aniline blue, connected to each other (Fig. 2G); in basal parts, cortex of one- or two-celled cortical layers, cells spherical or oval, and two to three layers of subcortical cells dichotomously concatenated at each layer, 52–55 μm thick (Fig. 2H), and a medulla of intertwined filaments with stellate cells (Fig. 2I & J); in distal part, cortex made up of one- or two-celled cortical layers, 5–7 × 8–11 μm, and one to two layers formed by subcortical stellate cells connected to anticlinal medullary filaments vertically (Fig. 2K), medulla most hollow and sparsely composed of anticlinal filaments, 5–7 μm wide, with medullary stellate cells, 17–18 × 18–20 μm (Fig. 2L); Tetrasporangia slightly ellipsoid to rounded, cruciately divided, 9–12 × 14–18 μm, scattered in the cortex (Fig. 2M). Gametophytes not found.
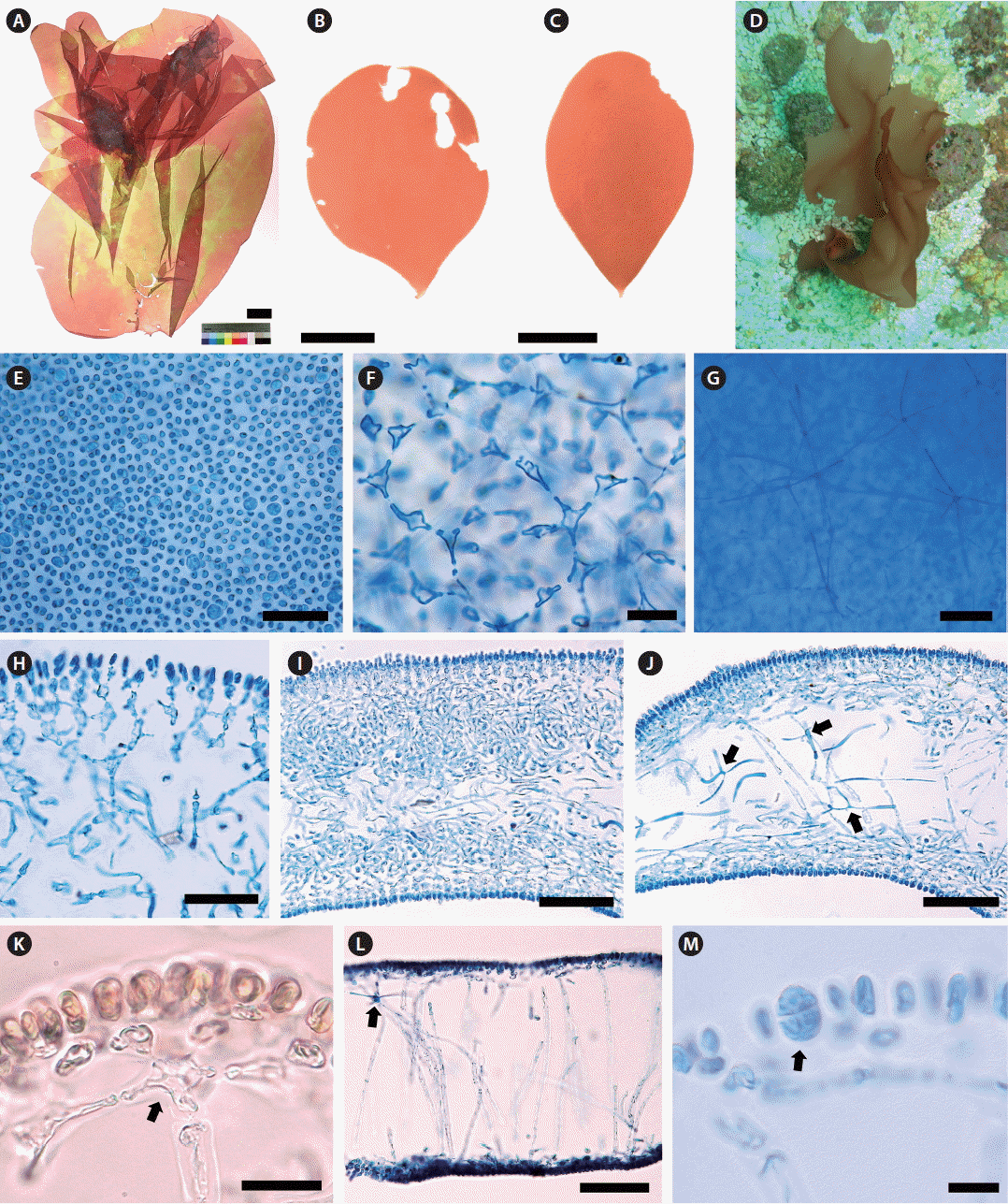
Amalthea rubida H. W. Lee et M. S. Kim sp. nov. (A) Holotype of A. rubida, a tetrasporophyte (MSK150624-42). (B & C) Habit of young plants MSK150531-07, vegetative, Dodu, Jeju, Korea on May 31, 2015 (B), MSK150620-04, vegetative, Dodu, Jeju, Korea on Jun 20, 2015 (C). (D) Habitat of A. rubida growing on a rhodolith bed at 17 m deep in Udo, Jeju, Korea. (E) Cortical cells in surface view. (F) Subcortical stellate cells connected to each other. (G) Projection of medullary stellate cells with long arms interconnected. (H) Cortex in the basal part. (I) Transverse section of the lowermost part nearby a stipe. (J) Transverse section of the basal part of thallus, arrows indicate medullary stellate cells with arms. (K) Cortex in the distal part, arrow indicates subcortical stellate cell. (L) Transverse section of the distal part of thallus, arrow indicates medullary stellate cells with arms. (M) Tetrasporangia in the cortical layer (arrow). Scale bars represent: A, 3 cm; B & C, 2 cm; E & H, 50 μm; F & K, 25 μm; G, I, J & L, 100 μm; M, 10 μm.
Holotype
MSK150624-42 (tetrasporophyte, KX879777, Fig. 2A), collected from 17 m deep of Udo, Jeju Island, Korea on Jun 24, 2015, and deposited in the Herbarium (JNUB), Jeju National University, Korea.
Isotype
MSK150624-40 (vegetative, KX879772, Udo, Jeju Island, Korea) deposited in JNUB and NIBRRD0000000184 (vegetative, MSK150624-41, KX879776, Udo, Jeju Island, Korea) deposited in the National Institute of Biological Resources (NIBR), Incheon, Korea.
Type locality
Udo, Jeju Island, Korea (33°30′27.75″ N, 126°56′08.49″ E).
Etymology
The specific epithet (rubida) was chosen to represent the color of this species when alive, being pale reddish brown to red.
Korean name
 .
.
Habitat
Amalthea rubida is growing in the subtidal habitat at a depth of 15–20 m attached to rhodoliths or pebbles, and the bottom of habitat is covered by sandy sediments with the strong tide current.
Other specimens examined
MSK150531-07 (vegetative, KX879773, Dodu, Jeju Island, Korea on May 31, 2015) (Fig. 2B); MSK150601-09 and MSK150601-10 (vegetative, Dodu, Jeju Island, Korea on Jun 1, 2015); MSK150620-04 (vegetative, KX879774, Dodu, Jeju Island, Korea on Jun 20, 2015) (Fig. 2C); MSK150721-01 (tetrasporophyte, KX879778, Dodu, Jeju Island, Korea on Jul 21, 2015); MSK150624-39 (KX879775), MSK150624-40 (KX879772) and NIBRRD0000000184 (MSK150624-41, KX879776) (vegetative, Udo, Jeju Island, Korea on Jun 24, 2015); MSK150801-02 (tetrasporophyte, KX879779, Udo, Jeju Island, Korea on Aug 1, 2015).
DISCUSSION
Our study of the molecular and morphological analyses of foliose red algae found a new species of Amalthea, and expanded the distribution for the genus, from New Zealand to Korea. The phylogenetic analyses of the rbcL gene supported the genus Amalthea and the recognition of our Korean sample as a new species, Amalthea rubida. Our results indicate that A. rubida and Halymenia abyssicola (GU598119) are sister taxa. Nevertheless, it is difficult to assign H. abyssicola to the genus Amalthea (D’Archino et al. 2014), because H. abyssicola from the Gulf of California is different only in thallus color (Dawson 1944, 1954) from A. freemaniae and A. rubida. It is necessary to carry out further morphological comparisons on generic criterion, such as reproductive structures of female specimens (D’Archino et al. 2014). Our data shows clearly that both species of Amalthea are distant from H. latifolia.
Based on the morphological characters of the generic type, A. rubida differs from A. freemaniae only in having a soft thallus with loosely arranged anticlinal filaments, and stellate cells (Table 1). There is also difference in the basal cortex thickness: six to eight layers in A. freemaniae (D’Archino et al. 2014), and three to five layers in A. rubida. These difference of cortex thickness can distinguish Amalthea species, like some Halymenia from the Gulf of California (Hernández-Kantun et al. 2009) and Pachymeniopsis from the northwestern Pacific (Yang and Kim 2015).
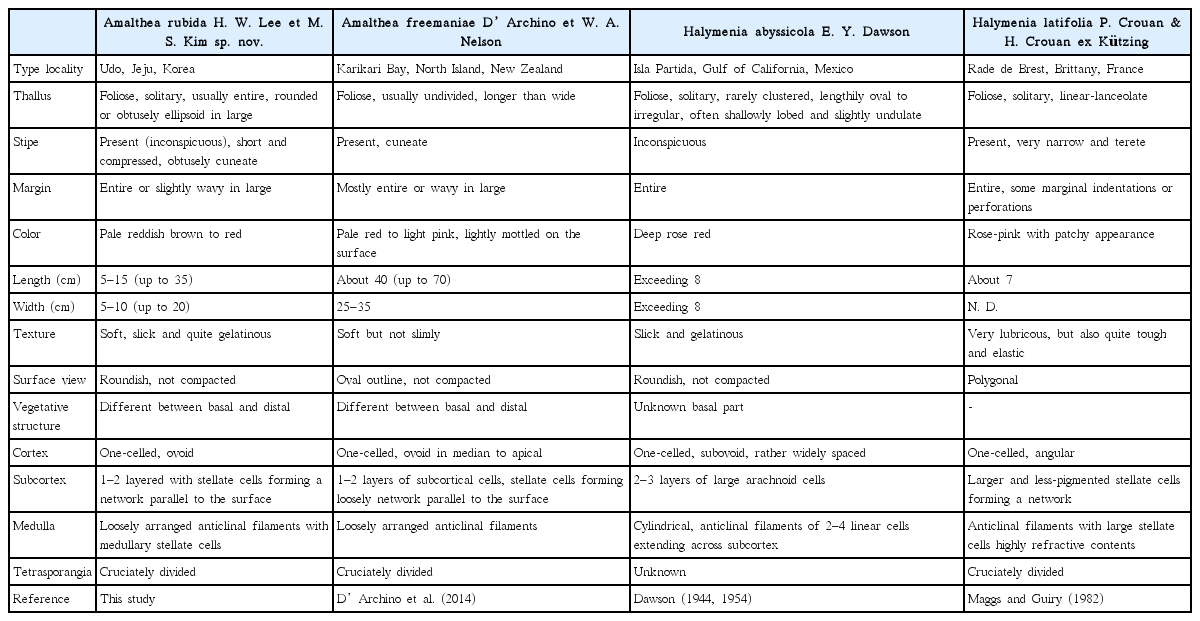
A comparison of taxonomic characteristics of Amalthea rubida with closely related taxa of the order Halymeniales
Amalthea rubida has not previously been found in Korea, possibly due to the difficulty accessing subtidal habitats at depths of 15–20 m. Among the order Halymeniales, Grateloupia kurogii Kawaguchi is similar (Kawaguchi 1990), and is found in the same subtidal habitat as A. rubida. However, mature G. kurogii has firm and yellow thalli consisting of a 4-celled outer cortex connected with a 3–4-celled inner cortex and periclinally or obliquely directed filaments in medullar with stellate cells (Kawaguchi 1990). Schizymenia apoda (J. Agardh) J. Agardh (Kim et al. 2012) may be indistinguishable from A. rubida in the subtidal regions only based on the thallus and texture. Anatomically, S. apoda is composed of anticlinal fascicles of dichotomously branched filaments in cortex and densely interwoven medullary filaments with hyphae (Gabriel et al. 2011, Kim et al. 2012). Galene rotunda is a little stiff, ellipsoid thallus composed of a 2–5-cell layered cortex and entwined medullary filaments that extend from a distinct stipe (Kim et al. 2012). Pachymeniopsis volvita is flat, circular to peltate, coriaceous thallus with ventrally curled margin, which consists of 6–9-cell layered cortex and intertwined filamentous medulla without distinctive stellate cells (Yang and Kim 2015).
The application of molecular data in the order Halymeniales has led to delimit the generic boundaries and species distribution (Gargiulo et al. 2013, D’Archino et al. 2014). Newly generated sequences of the rbcL gene show the sufficient interspecific divergence to determine species diversity, for example, Cryptonemia asiatica (Yang and Kim 2014) and Pachymeniopsis volvita (Yang and Kim 2015). In the genus Amalthea, D’Archino et al. (2014) mentioned that molecular data can only identify this genus because its morphological characteristics are not exclusive. Our rbcL sequences of Amalthea rubida from eight samples in Korea were identical and formed a clade with the type species A. freemaniae (KJ606651-KJ606653) from New Zealand and H. abyssicola (GU598119) from the Pacific Mexico (Azevedo et al. 2016). The vegetative difference between A. rubida and A. freemaniae in the basal cortex thickness was well reflected in the rbcL phylogenetic tree. The recognition of A. rubida from Korea enabled us to separate several species such as Grateloupia kurogii and Schizymenia apoda having very similar morphology (Kim et al. 2012, Yang and Kim 2015).
Recently, the endemism of the temperate southwestern Pacific region was mirrored in the establishment of new genera from New Zealand: Amalthea (D’Archino et al. 2014), Judithia, Wendya (D’Archino et al. 2016), Psaromenia (D’Archino et al. 2010), and Stauromenia (D’Archino et al. 2012). The global endeavor to resolve misunderstood or misidentified specimens lead to recognize a new distribution of the genus known as endemic. Like recognizing a new distribution of Psaromenia (Schneider et al. 2014), our study revealed an expanded distribution of the genus Amalthea by discovering A. rubida. The distribution of Amalthea and Psaromenia in both hemispheres and crossing the tropical Pacific makes it difficult to fully understand how this distribution came about. Adey and Steneck (2001) proposed that in the western Pacific, several large intercontinental seas, coastal water was less affected by the continental climate and more directly affected by ocean currents during glacial times on the basis of the thermographic model. Further collection, especially in the tropical western Pacific where Amalthea may exist, may clarify how this dispersal could have occurred.
This study describes a new species of Amalthea, which was originally a monospecific and endemic genus in New Zealand, and expands the distribution of from the southern hemisphere into the northern hemisphere. The accumulation of molecular data with morphological observations provides valuable opportunities to define the taxonomic position of foliose red algae accurately, and to understand algal distribution patterns.
ACKNOWLEDGEMENTS
We thank all members of the molecular phylogeny of marine algae laboratory at Jeju National University. This work was supported by a grant from the National Institute of Biological Resources (NIBR), funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR201624202 and NIBR201501204).