ABSTRACT
Dolichospermum is a filamentous and heterocytous cyanobacterium that is one of the commonly occurring phytoplanktons in the Han River of Korea. Morphological observations led to the identification of D. planctonicum-like filaments in seasonal water samples. In the present study, we successfully isolated these filaments using culture methods, and examined its morphology using light and scanning electron microscopy. The morphology of the D. planctonicum-like species differed from that of typical D. planctonicum; it had thin cylindrical-shaped akinetes, which were narrower towards the ends than at the center. This morphology is firstly described in the genus Dolichospermum. In addition, the akinetes in the filament developed solitarily and were distant from the heterocytes. Phylogenetic analysis of the 16S rRNA sequences showed that our Dolichospermum clustered with D. planctonicum and D. circinale, which have coiled trichome. However, phylogenetic analysis of the gene encoding rivulose-1,5-bisphosphate carboxylase (rbcLX) clearly separated our species from other Dolichospermum, forming a unique clade. Additionally, structures of D. planctonicum and D. hangangense strains were different type in Box-B and V3 region. These results demonstrated that the new Dolichospermum species was unique in morphology and molecular traits. Therefore, we propose this to be a new species belonging to genus Dolichospermum with the name Dolichospermum hangangense sp. nov.
INTRODUCTIONThe cyanobacterium Dolichospermum occurs in freshwater bodies worldwide, although they mostly dominate rivers or lakes in the summer season (Oliver and Ganf 2000, Zhang et al. 2016). Some species (e.g., Dolichospermum circinale, D. planctonicum, D. smithii, and D. spiroides) of this genus generate harmful algal blooms, which deteriorates water quality and adversely affects aquatic environments.
From the historical perspective, Bornet and Flahault (1888) first named this genus Anabaena. However, Wacklin et al. (2009) suggested that planktic and benthic mat-forming Anabaena can be classified using 16S rRNA phylogenetic analysis. A recent study showed that planktonic Anabaena cells containing gas vesicles could be newly classified as genus Dolichospermum (Wacklin et al. 2009), whereas benthic mat-forming Anabaena characterized the genus Anabaena (Bornet and Flahault 1888). Hence, the first holotype species of Dolichospermum was named Anabaena flos-aquae (recently it was renamed to Dolichospermum flosaquae) (Brébisson ex Bornet and Flahault 1888) (see Wacklin et al. 2009). Currently, 55 species have been described taxonomically in the genus Dolichospermum after considering the AlgaeBase database (Guiry and Guiry 2013).
Overall, the genus Dolichospermum is characterized by morphological characteristics, such as types of trichomes, length and width, shape, position of akinetes, morphology of vegetative cells and heterocytes, and existence of mucilaginous sheath, etc. (Mishra et al. 2015). However, the species-specific characteristics may show phenotypic overlap with features of other species. In addition, these features are transformed by different environmental conditions and long culture periods (Rippka et al. 1979, Watanabe et al. 2004). Additionally, it is difficult to determine the key identifying morphological characters in field samples. Hence, Watanabe et al. (2004) suggested considering maturation of akinetes for circumventing the fundamental limitations regarding morphology-based identification. A recent study showed that morphological variations of the akinetes occurred during various maturation steps in the genus Cylindrospermum (Nostocales) (Johansen et al. 2014).
Phylogenetic analysis using 16S rRNA (small-subunit rRNA-encoding DNA) was performed to corroborate the results of morphology-based classification (Lee et al. 1996, Zapomělová et al. 2008). However, the position of species or genus of filamentous cyanobacteria, including those in the order Nostocales, in the 16S rRNA-based phylogenetic tree was ambiguous (Wilmotte and Herdman 2001, Gugger et al. 2002, Gugger and Hoffmann 2004). Therefore, the 16S rRNA was not sufficient for supporting the results of morphology-based identification. As an alternative solution, Singh et al. (2015) suggested that the neutrally evolving genes rbcL (encoding D-ribulose 1,5-bisphosphate carboxylase-oxygenase large subunit) and the less conserved rbcX were appropriate for classifying cyanobacteria. Additionally, an internally transcribed spacer between 16S rRNA and 23S rRNA genes (16S-23S ITS) have more variable regions and can separate morphologically similar species (Li and Brand 2007). Therefore, analyses of rivulose-1,5-bisphosphate carboxylase (rbcLX) and 16S-23S ITS genes could support to taxonomically evaluation.
The Han River is the most important water resource in South Korea, because it runs through the center of Seoul, which is inhabited by a quarter of the whole Korean population. In addition, it serves as a valuable source of water for drinking and irrigation (Han et al. 1993, 1995, Joo et al. 2016b, 2017). Thus, continuous monitoring in the Han River is important to management of water quality (Han et al. 1999, 2002, Chang 2005, Suh et al. 2005). However, cyanobacterial bloom occurs in the river and reservoir, which is caused mainly by genus Dolichospermum, and can cause ecological and economic problems (Kim et al. 2005, Li et al. 2015, Joo et al. 2016a). Morphological observations showed the presence of D. planctonicum-like cells several times in the Han River. However, morphological observations could not uncover the species identity of this organism of genus Dolichospermum (Li et al. 2013).
In the present study, we isolated D. planctonicum-like cells using culture methods in the laboratory, and examined its morphology using light and scanning electron microscopy. We characterized the surface morphology of our strain and compared it with those of four closely related species (D. circinale, D. planctonicum, D. smithii, and D. spiroides). In addition, molecular analyses were performed with 16S rRNA, rbcLX, and 16S-23S ITS sequences.
MATERIALS AND METHODSWater samples were collected from lakes Soyang (38°00′31.76″ N, 127°54′21.60″ E), Chuncheon (37°58′41.79″ N, 127°40′16.45″ E), Uiam (37°51′43.27″ N, 127°41′16.03″ E), and Cheongpyeong (37°42′31.42″ N, 127°27′07.30″ E) situated on the Han River, and the river located in Seoul, South Korea in 2013. We isolated single trichomes from the samples using a Pasteur pipette under an inverted microscope (Olympus IX, Tokyo, Japan), and the isolated cells were grown in 1 mL CB medium (pH 9.0) (Ichimura and Itoh 1977) and incubated in 48-well plates. In this study, we successfully cultured four strains of Dolichospermum, including D. circinale, D. smithii, D. planctonicum, and D. spiroides, from the Han River samples. They were maintained under constant culture conditions (20°C, 45 μmol photons m−2 s−1 light intensity, 12: 12 h light: dark cycle). The surviving cells were transferred to culture flasks. Morphological characteristics of these four Dolichospermum species were evaluated using fresh samples. A portion of each sample was preserved with Lugol’s solution (2.5%), and the remainder was used for cell isolation.
The Dolichospermum strains cultured in this study were examined using a BX 51 photomicroscope (Olympus) and cell dimensions were measured using the Cellsens program v.1.8 (Olympus). The following morphological characteristics were determined for at least 30 cells: length and width of vegetative cells and akinetes, and coil diameter and distance between coils for coiled trichomes. Based on all morphometric features, species groupings were performed using analysis of variance (ANOVA) of the SPSS Statistics v.17.0 (SPSS Inc., Chicago, IL, USA). A p-value < 0.05 was considered significant by Duncan’s test.
For more detailed observation using a scanning electron microscopy (SEM), cells obtained from growing cultures were fixed in 2.0% glutaraldehyde solution (Sigma-Aldrich, St. Louis, MO, USA) and stored at 4°C for 24 h. The fixed samples were rinsed with distilled water to completely remove all fixation reagents and dehydrated for 10 min each with series of ethanol treatments (30, 50, 70, 90, and 95% ethanol). Finally, the samples were dehydrated twice for 20 min with 100% ethanol. The dehydrated samples were dried using critical point dryer (CPD 030; BAL-TEC, Walluf, Germany), coated with gold for 150 s using 40 mA currents (BAL-TEC SCD 005 super coater; BAL-TEC, Liechtenstein, Germany), and examined using a SEM (NOVA NANO SEM 450; FEI, Eindhoven, Netherlands).
The biomass of the studied strains was harvested at the exponential growth stage. Genomic DNA was extracted using a DNeasy plant kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. The polymerase chain reaction (PCR) mixture (total volume 20 μL) consisted of 11.9 μL sterile water, 2 μL 10× Taq buffer, 2 μL dNTP, 0.1 μL Ex Taq of the Takara Ex Taq PCR kit (Takara Shuzo, Osaka, Japan), 1 μL of each primer (0.01 mM), and 2 μL template DNA. PCR was performed on an icycler (Bio-Rad, Hercules, CA, USA) using 16S rRNA primers CYA108F (5′-ACG GGT GAG TAA CRC GTR A-3′) (Nübel et al. 1997) and 16S CYR (5′-CTT CAY GYA GGC GAG TTG CAG C-3′) (Urbach et al. 1992) and amplified using the following steps: initial denaturation at 94°C for 5 min, followed by 30 cycles of 94°C for 1 min, 57°C for 1 min, and 72°C for 1 min, and a final extension step of 72°C for 5 min.
PCR amplification of rbcLX was performed with primers CW (5′-CGT AHC TTC CGG TGG TAT CCA CG T-3′) and CX (5′-GGG GCA GGT AAG AAA GGG TTT CGT A-3′) (Rudi et al. 1998). In addition, amplification of rbcLX was performed in two steps as follows: initial denaturation of the first step at 94°C for 4 min, two cycles of 94°C for 30 s, 40°C for 30 s, 72°C for 2 min, followed by initial denaturation of the second step at 94°C for 30 s, followed by 38 cycles of 55°C for 30 s, 72°C for 2 min, and a final extension reaction at 72°C for 7 min.
Phylogenetic analyses were performed with the sequences of our strain and other sequences of the same genus retrieved from the public database. They were aligned using the BioEdit program v.7.1.3 (Hall 1999). The alignment was edited manually. We determined partial sequences of the 16S rRNA (1,277 bp) and rbcLX (787 bp). Sequences were deposited in GenBank (http://www.ncbi.nlm.nih.gov) with accession numbers KR154298-KR154317 (Table 1).
The topology for the phylogenetic tree was derived using the maximum likelihood (ML) method. The GTR + G + I (the General Time Reversible model incorporating in variant sites and a gamma distribution) model was selected for Bayesian and ML analyses as determined by MrModelTest v.2.3 (Nylander 2004). In Bayesian analysis, the Markov chain Monte Carlo process was set at four chains and 10,000,000 generations. The sampling frequency was 100 generations. The first 25% of all trees were deleted as burn-ins, and a consensus tree was constructed in MrBayes v.3.2.3 (Ronquist and Huelsenbeck 2003). The MEGA 7.0 software was used for ML and neighbor joining phylogenetic analysis using the p-distance models. All analyses were tested using bootstrapping with 1,000 replicates.
Additionally, PCR amplification of 16S-23S ITS was performed with primers 322 (5′-TGT ACA CAC CGC CCG TC-3′) and 340 (5′-CTC TGT GTG CCT AGG TAT CC-3′) (Laloui et al. 2002). Amplification of 16S-23S ITS was performed in two steps as follows: initial denaturation of the first step at 94°C for 10 min, 38 cycles of 94°C for 30 s, 55°C for 30 s, 72°C for 1 min and a final extension reaction at 72°C for 7 min. The Box-B and V3 helices of ITS edited in BioEdit program v.7.1.3 (Hall 1999). The secondary structures were constructed using Mfold 3.1 (Zuker 2003).
RESULTSDolichospermum hangangense H. J. Choi & M. -S. Han sp. novDiagnosisTrichomes solitary, straight filaments; vegetative cells barrel-shaped to spherical, 5.1–11.5 μm wide and 5.0–10.1 μm length, length: width (L: W) ratio 0.5–1.8, with gas vesicles. Heterocytes round, colorless or light green. Akinetes light green, thin cylindrical, narrower towards the ends than at the center, 10.9–15.8 μm wide and 19.7–51.9 μm length, L: W ratio 1.7–3.8, separate from heterocytes.
NeotypeFixed sample in 2.5% Lugol’s solution of strain HYCP1307-DH5 and HYCC1307-DH38 collected from the Han River in July 2013 were stored in the laboratory for water environmental ecology restoration, Department of Life Science, Hanyang University, Seoul, South Korea.
Type localityLakes Chuncheon (37°58′41.79″ N, 127°40′16.45″ E) and Cheongpyeong (37°42′31.42″ N, 127°27′07.30″ E), Han River, Seoul, South Korea.
Etymology‘Dolichospermum hangangense.’ ‘hangang’ derived from korean name of Han River and ends of name ‘-ense’ was considered based on formation of bacterial names from geographical names in Bergey’s manual of systematic bacteriology (Garrity et al. 2012).
Morphological characterization of Dolichospermum hangangenseWe observed the morphology of new species. The akinetes were emerald green in color and had thick mucilaginous sheath. Thin cylindrical akinetes were narrower towards the ends than at the center (Fig. 1A). The surface of the akinetes was observed by using SEM. Mucilaginous sheath enveloped to surface of akinetes. The surface of akinetes was smooth and bandless (Fig. 1B). The width and length of akinetes were 10.9–15.8 × 19.7–51.9 μm and the L: W ratio of the akinetes was 1.7–3.8. Positions of akinetes were always solitary, separate from heterocytes (Table 1). The thicken mucilaginous sheath formed on either side of the akinetes resembled a fingernail when akinetes were separated from trichomes. The trichomes were straight and solitary, and were not in fascicles. The vegetative cells were rounded or barrel-shaped along with cell division stages and contained gas vesicles. The cell was dark green in color (Fig. 1C). The width and length of vegetative cells were 5.1–11.5 × 5.0–10.1 μm and the L: W ratio of the vegetative cells was 0.5–1.8 (Table 1). Distinct characteristics of this new species appeared during maturation stages, when rounded mucilaginous envelopes covered the akinetes. The sheath, which was colorless and distinct, formed only in immature akinetes of this species (Fig. 1D). The heterocytes were almost round and colorless or light green (Fig. 1E). Additionally, we observed the morphological variations of akinetes among species (D. circinale, D. planctonicum, D. smithii, and D. spiroides) related to D. hangangense sp. nov. in maturation stages. In immature stage, the akinetes of all species had incomplete morphology. The immature akinetes had light green color that was similar to that of vegetative cells; cell size was larger than those of vegetative cells, and they were round or slightly oval in shape (Fig. 2A–D). The matured akinetes of the four species were elongated and wider than the immature akinetes. The ripen akinetes of D. circinale were spherical and slightly ellipsoidal, and their width and length ranged from 16.6–22.7 μm and 17.4–32.0 μm, respectively; the L: W ratio was 0.8–1.5. The akinetes were present as solitary bodies, away from heterocytes (Table 1, Fig. 2E). The ripen akinetes of D. smithii were spherical, and their width and length ranged from 15.4–28.3 μm and 18.8–30.8 μm, respectively; the L: W ratio was 1.0–1.6. The akinetes were present as solitary bodies, away from heterocytes (Table 1, Fig. 2F). The ripen akinetes of D. planctonicum were ellipsoidal or cylindrical. Their width and length ranges were 12.8–25.0 μm and 13.8–45.0 μm, respectively; the L: W ratio was 1.1–2.6. They were present as solitary bodies or were nearby heterocytes (Table 1, Fig. 2G). Finally, the ripen akinetes of D. spiroides were mostly ellipsoidal, with width and length ranging from 9.0–12.0 μm and 13.6–21.1 μm, respectively. The L: W ratio of the akinetes was 1.4–2.0, and they occurred as solitary structures distant from heterocytes (Table 1, Fig. 2H). The morphological characteristics and morphometric features of the akinetes of D. hangangense and its relatives revealed by phylogenetic analysis are summarized for inter-species comparison and better understanding of their relatedness (Fig. 3).
Molecular phylogeny of Dolichospermum hangangensePhylogenetic analyses using a broad range of freshwater strains including genus Dolichospermum based on morphological characteristics confirmed with previous studies. We evaluated the 16S rRNA sequences (1,277 bp) of D. hangangense (HYCP1307-DH5 and HYCC1307-DH38) using sequences of its phylogenetic relatives. These strains were grouped together with other species including genus Dolichospermum in phylogenetic tree of 16S rRNA. The closest relative was D. circinale 04–29 (100% coverage and 99% DNA identity). The secondary close relatives were Dolichospermum sp. TAC472 having same morphology of new species (98% coverage and 99% identity) and Dolichospermum planctonicum NIES 817 and 1,681 (92% coverage and 99% identity). The sequence of D. hangangense was proximal to those of its closest relatives in the 16S rRNA-based phylogenetic tree. However, there is no close relatives which were totally matched the sequences. In phylogenetic tree of 16S rRNA, D. hangangense were closest to a species D. circinale. These species have coiled trichome and almost spherical and oval akinetes. The secondary relatives were D. planctonicum. This species has straight trichome and oval akinetes (Fig. 4).
In addition, we used the sequence of rbcLX (787 bp) from D. hangangense for comparison with those of its relatives. In phylogenetic tree of rbcLX, D. hangangense strains made a solitary branch with only Dolichospermum sp. TAC 472 strain contrary to the phylogenetic results of 16S rRNA. This strain having the same morphological characteristics with new species was showed the highest sequence similarity with D. hangangense strains (97% coverage and 99% identity). A phylogenetically close taxon was composed of D. circinale and D. planctonicum group strains like a phylogenetic result of 16S rRNA (Fig. 5).
Secondary structureWe conducted secondary structure analysis to distinguish between the new speices and D. planctonicum having most similar morphological characteristics and sequences in 16S-23S ITS gene to new species. Box-B and V3 region of 16S-23S ITS gene were commonly used to distinguish species of cyanobacteria (Li and Brand 2007). D. hangangense was identical in secondary structure of the Box-B helix, V3 helix to these 11 strains (eight D. planctonicum, two D. hangangense, and one Dolichospermum sp. strains). In Box-B region, all of D. planctonicum strains were having the same structures. Structure of D. planctonicum strains had three loops. The total sequences of Box-B region consisted of 48 bases. Structure of D. hangangense strains had 4 loops. Total structure sequences of HYCP1307-DH5 and HYCC1307-DH38 strains consisted of 56 and 58 bases. However, structure of D. hangangense were having some variable part within the species. The sequences of the middle of Box-B helix differed in sequences each strain (GAAGU vs. GAAGUC and CCUA vs. GCCUC). Additionally, structure of TAC472 strain was having same morphological characteristics with higher variablity than structure of D. hangangense strains. The total structure sequences of TAC 472 consisted of 51 bases (Fig. 6). Box-B region was having higher variable parts than V3 region. In V3 region, D. planctonicum strains also had same structures. Structure of D. planctonicum strains had 3 loops. The total sequences of D. planctonicum strains consisted of 38 bases. Structure of D. hangangense strains had three loops. The total sequences of D. hangangense strains consisted 48 bases. However, strains of D. hangangense had some variable part within the species. In addition, structure of TAC472 strain in V3 region was also having same morphological characteristics with higher variablity than structure of D. hangangense strains. Structure of TAC472 had four loops. The total sequences of TAC472 strain consisted of 45 bases (see Fig. 7).
DISCUSSIONIn taxonomy, species of the genus Dolichospermum are classified based on morphology. However, this system of classification has its limitations. Most trichomes isolated from field samples of lakes and/or rivers lack akinetes as the key characteristics (Singh 1973, Fiore et al. 2005, Johansen et al. 2014). In other words, only few taxa without akinetes could be distinguished based on the morphological characteristics of the vegetative cell, heterocytes, end cell, mucilaginous sheath surrounding trichomes. Even filamentous cyanobacteria, including those in the genus Cylindrospermum, showed that morphological characteristics changed during maturation (Johansen et al. 2014). Johansen et al. (2014) showed that the akinetes of Cylindrospermum alatosporum changed from round cylindrical to thin cylindrical shape during maturation. The matured akinetes of C. catenatum developed a brown color. In the present study, we examined the morphological characteristics of four Dolichospermum species during maturation (see Fig. 2A–D). The morphology of D. hangangense was like that of immature D. planctonicum (Figs 1C & 2C). However, during aging, the akinetes of two species showed distinguishable key characteristics. The morphology of the akinetes of D. planctonicum did not change (Fig. 2G). In contrast, the akinetes of D. hangangense were narrower towards the ends than at the center, and rounded mucilaginous sheath surrounded only the akinetes during maturation (Fig. 1D). Finally, the ripen akinetess of D. hangangense had thin cylindrical with higher L: W ratio (2.3–2.6) in morphometric comparisons with D. planctonicum (see Table 1). Therefore, we propose that we should use totally ripen akinetes for precise identification of akinetes-forming cyanobacteria. Additionally, we could distinguish morphologically similar species, based on their morphological changes during their maturation stages.
We also conducted phylogenetic analysis of the 16S rRNA with several complete and partial sequences of cyanobacterial strains deposited in the National Center of Biotechnology Information (NCBI) database. The high similarity in 16S rRNA sequences among diverse species of Dolichospermum supported the features of the morphology-based phylogenetic tree. Particularly, sequence similarity between D. planctonicum and D. hangangense was 99.5–99.7%. Similarly, sequence similarity between D. planctonicum and D. hangangense of rbcLX was 98.5–98.6%. In other words, phylogenetic analysis of rbcLX is preferred for classifying species than that based on 16S rRNA sequence. Singh et al. (2015) suggested that phylogenetic analyses using the sequence of rbcLX as a molecular marker for classifying species, including those in the order Nostocales, can be used for precise identification of heterocytous cyanobacteria with ambiguous classification. In the present study, we observed that the 16S rRNA sequence was incongruent as a criterion for distinguishing taxonomical diversity in the order Nostocales (Johansen et al. 2014). In addition, molecular analyses of the two studied genes did not solve the problem regarding classification of heterocytous cyanobacteria. Typical cut-offs for taxon recognition of cyanobacteria (<95% for genus and <97.5% for species) (Stackebrandt and Goebel 1994, Ludwig et al. 1998) were conservative for classification of cyanobacteria (Lukešová et al. 2009, Vaccarino and Johansen 2012). In other words, these cut-offs were suitable for only certain species (Komárek and Mareş 2012). Therefore, cut-offs should more tight and closer than current standards for exact classification of cyanobacteria. Interestingly, we observed that the TAC 472 strain from Japan was closer to D. planctonicum in phylogenetic analysis. However, this strain formed the same clade with D. hangangense as observed with the 16S rRNA and rbcLX phylogenetic analyses (Figs 4 & 5). In addition, we also confirmed the morphological characteristics and morphometric features of the strains, which showed that this strain was identical to D. hangangense sp. nov. Therefore, the sequence of TAC 472 deposited in NCBI should be considered for re-identification of species for accurate classification. Additionally, we conducted secondary structure analysis for distinguishing between new species and D. planctonicum having most similar morphological characteristics and sequences in 16S–23S ITS gene. Box-B and V3 helices were the most variable in heterocytous cyanobacteria, 16S–23S ITS secondary structures can be promising markers to be discriminated at the intra-generic level of the genus Dolichospermum (Li and Brand 2007). Structures of D. planctonicum and D. hangangense strains were different type in Box-B and V3 region. In addition, we found that structures of D. hangangense strains had some variable sequences and structures within the species (Figs 6 & 7).
The current taxonomy of cyanobacteria still depends extensively upon morphological characteristics and molecular analysis of the ribosomal RNA-encoding gene. However, these methods are not enough for the classification of various species, including those in the order Nostocales, of cyanobacteria. Thus, analysis of polyphasic genes related to biochemical (cyanotoxins, oligopeptides, etc.) (Pearson et al. 2010) and ecophysiological characters and others (rpoB, mcy-genes, gvp-genes, and nif-genes) (Rajaniemi et al. 2005, Tanabe et al. 2007) should be considered for exact classification (Komárek and Mareş 2012).
In conclusion, we reported that novel Dolichospermum species, Dolichospermum hangangense was taxonomically described based on morphological and molecular analysis.
ACKNOWLEDGEMENTSThis research was supported by the Environmental Basic Research Program, Han River watershed management committee.
Fig. 1Morphological characteristics of Dolichospermum hangangense sp. nov. (HYCP1307-DH5) isolated from the Han River. (A) Ripen akinetes in trichome. The arrows mark the curved line. Akinete shape was narrower towards the ends than at the center. (B) Scanning electron microscopy showing the surface morphology of the akinete of D. hangangense. (C) Trichome of D. hangangense. Distinct mucilaginous sheath was formed surrounding the akinetes. The thickened mucilaginous sheath resembled a fingernail when akinetes were separated from the trichome. (D) Immature akinetes surrounding round mucilaginous sheath. This akinete had incomplete morphology and was narrower towards the ends than at the center. (E) Vegetative cells in the trichome. Vegetative cells have different shape, from round to barrel-shaped in various cell cycle phases. Scale bars represent: A–E, 10 μm. 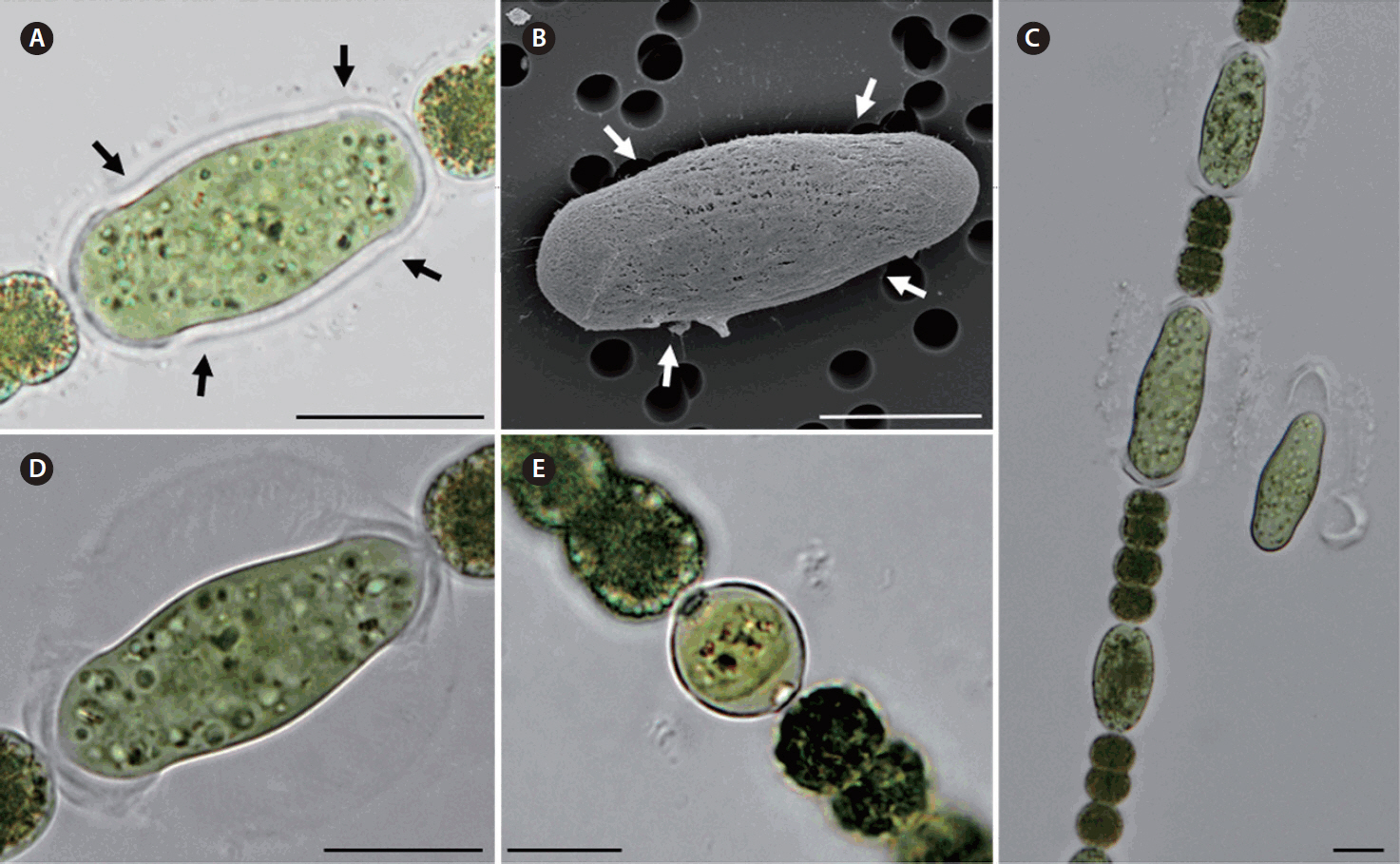
Fig. 2Light micrographs of the species proximal to Dolichospermum hangangense sp. nov. in the 16S rRNA phylogenetic tree showing morphological characteristics of immature akinetes (A–D). (A) D. circinale. (B) D. smithii. (C) D. planctonicum. (D) D. spiroides. Light micrographs of species proximal to D. hangangense sp. nov. in the 16S rRNA phylogenetic tree showing morphological characteristics of matured akinetes (E–H). (E) D. circinale. (F) D. smithii. (G) D. planctonicum. (H) D. spiroides. Scale bars represent: A–H, 10 μm. 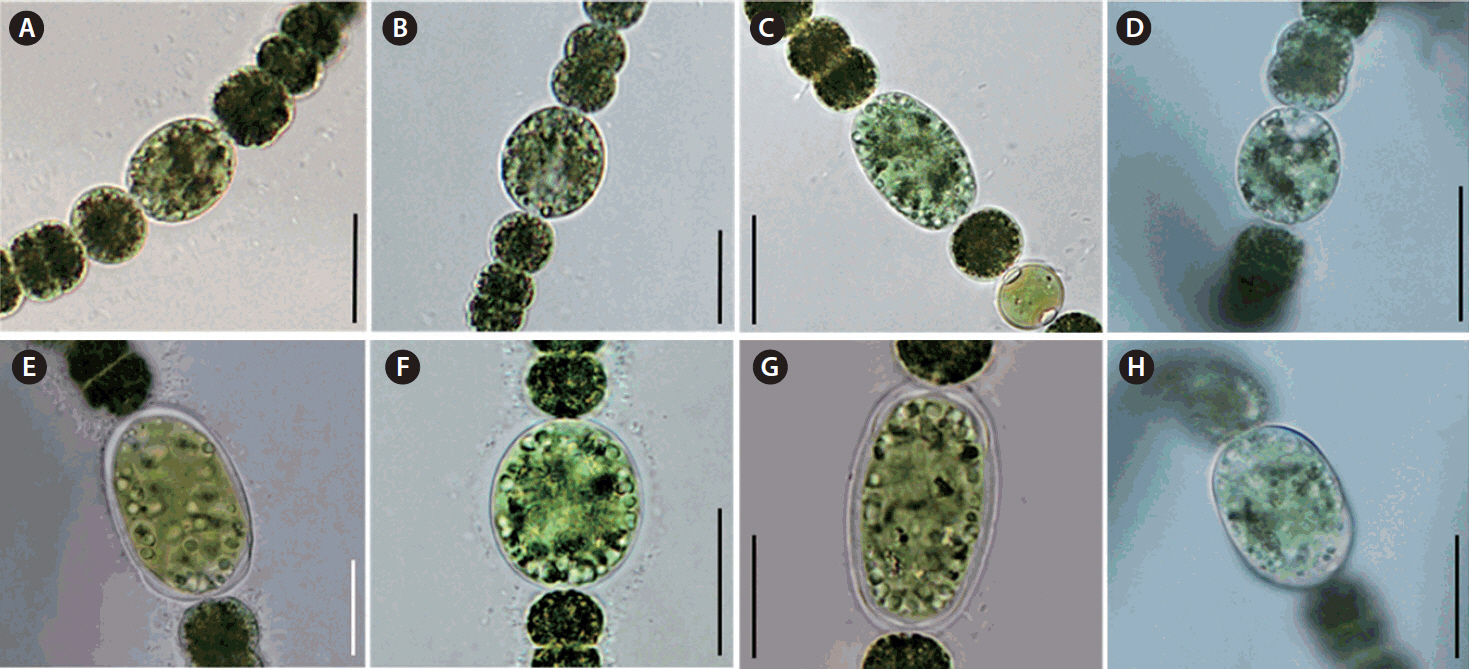
Fig. 3A drawing showing the morphological characteristics of ripen akinetes of Dolichospermum hangangense sp. nov. and its neighboring species in the 16S rRNA phylogenetic tree. *The length: width ratios of akinetes of various Dolichospermum species. ‘n’ indicate the number of akinetes for measuring length and width. 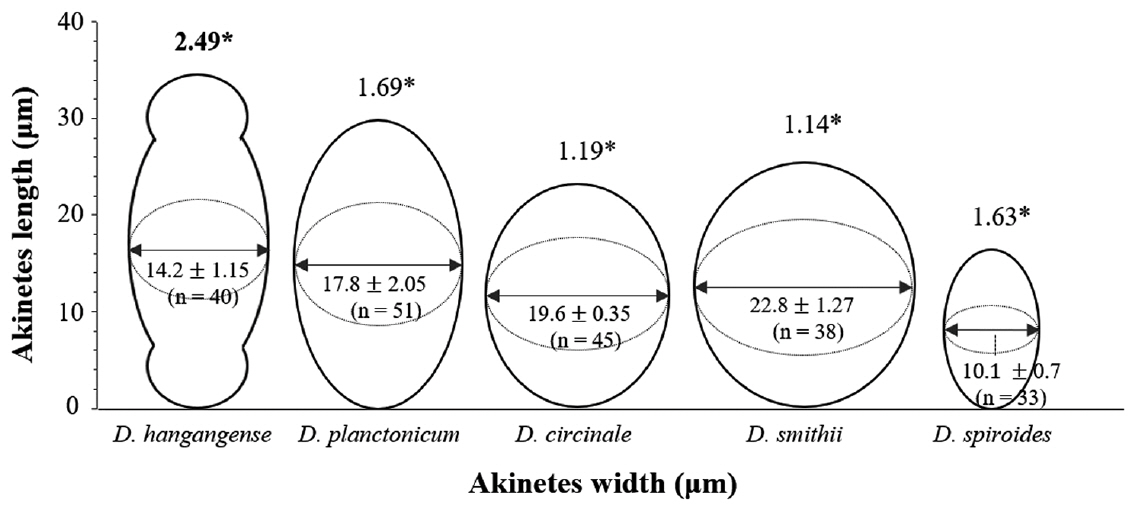
Fig. 4Neighbor joining based on 16S rRNA (1,277 bp) sequences showing the clustering of cultured Dolichospermum spp. strains. #Tentatively assigned name, because its species checked morphological characteristics. This species had same morphology with D. hangangense. Numbers above branches indicate bootstrap support (>50%) from neighbor joining/maximum likelihood/Bayesian analyses. D. hangangense has gray background. Calothrix sp. PCC 6303 was used as the out group. 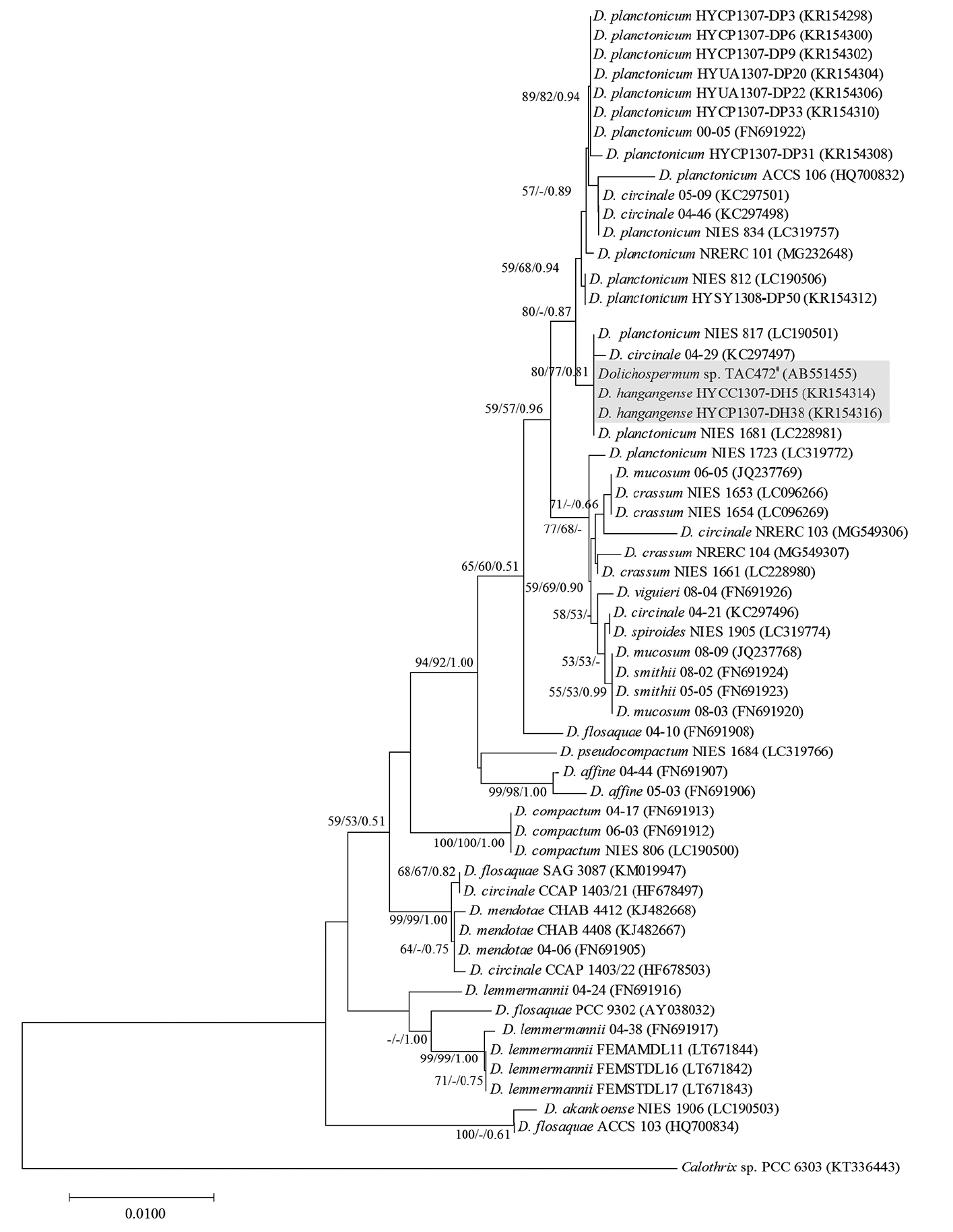
Fig. 5Neighbor joining based on rbcLX (787bp) sequence showing the clustering of cultured Dolichospermum spp. strains. #Tentatively assigned name, because its species checked morphological characteristics. This species had same morphology with D. hangangense. Numbers above branches indicate bootstrap support (>50%) from neighbor joining/maximum likelihood/Bayesian analyses. D. hangangense has gray background. Nostoc punctiforme SAG 69.79 was used as the out group. 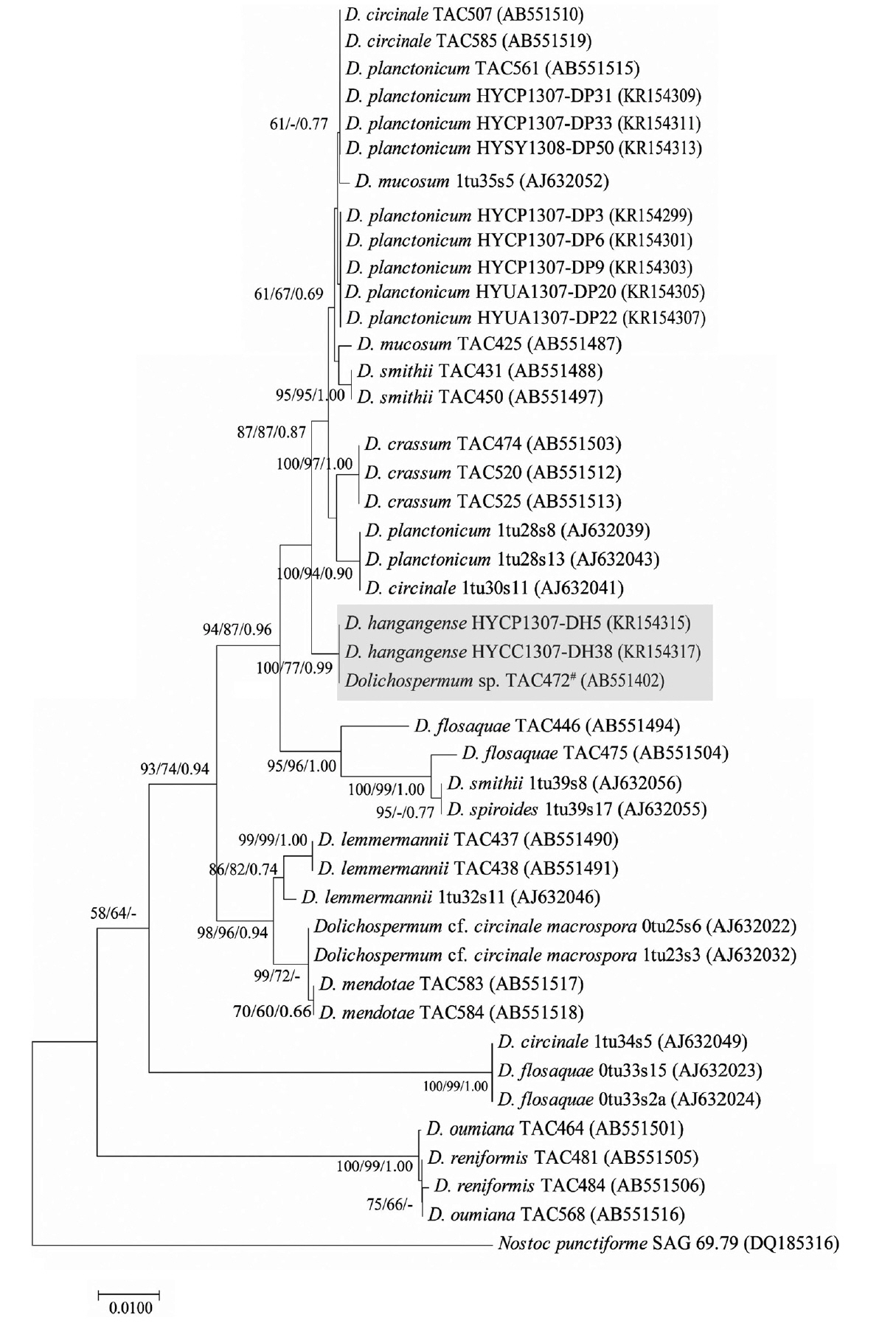
Fig. 6Secondary structures of the Box-B helix in Dolichospermum spp. strains. Arrows and gray background show bases variable within the new species. 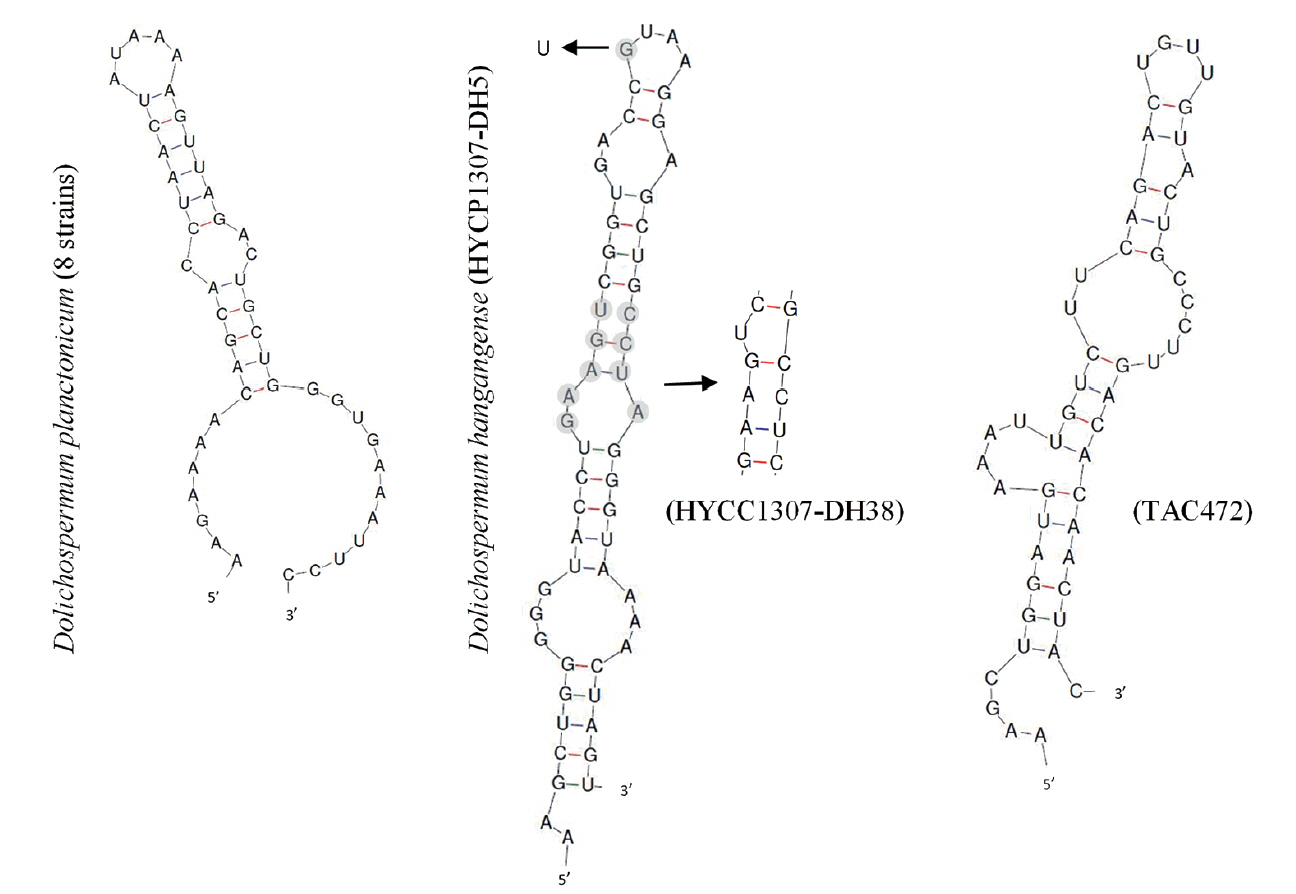
Fig. 7Secondary structures of the V3 helix in Dolichospermum spp. strains. Arrows and gray background show bases variable within the new species. 
Table 1Morphometric features of the studied cyanobacterial strains used in this study
REFERENCESBornet, É. & Flahault, C. 1888. Révision des Nostocacées hétérocystées contenues dans les principaux herbiers de France (quatrième et dernier fragment). Annales des Sciences Naturelles, Botanique, Septième, Série. 7:177–262.
Chang, H. 2005. Spatial and temporal variations of water quality in the Han River and its tributaries, Seoul, Korea, 1993–2002. Water Air Soil Pollut. 161:267–284.
Fiore, MF., Nelian, BA., Copp, JN., Rodrigues, JLM., Tsai, SM., Lee, H. & Trevors, JT. 2005. Characterization of nitrogen-fixing cyanobacteria in the Brazilian amazon floodplain. Water Res. 390:5017–5026.
Garrity, G., Boone, DR. & Castenholz, RW. 2012. Bergey’s manual of systematic bacteriology. Vol. 1. The Archaea and the deeply branching and phototrophic bacteria. Springer-Verlag, New York, 722 pp.
Gugger, M., Lyra, C., Henriksen, P., Couté, A., Humbert, JF. & Sivonen, K. 2002. Phylogenetic comparison of the cyanobacterial genera Anabaena and Aphanizomenon. Int. J Syst Evol Microbiol. 52:1867–1880.
Gugger, MF. & Hoffmann, L. 2004. Polyphyly of true branching cyanobacteria (Stigonematales). Int J Syst Evol Microbiol. 54:349–357.
Guiry, MD. & Guiry, GM. 2013. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway, Available from: http://www.algaebase.org. Accessed Aug 1 2017
Han, M. -S.., Auh, Y. -Y.., Ryu, J. -K.., Yoo, K. -I.. & Choi, Y. -K. 1995. Ecological studies on Pal’tang River-Reservoir System in Korea 2. Changes in phytoplankton community structure. Korean J Limnol. 28:335–344.
Han, M. -S.., Kim, Y. -O.., Yi, D. -S.. & Hong, S. -S. 2002. Species-specific productivity of Cyptomonas ovata (Cryptophyceae) in the Pal’tang Reservoir, Korea. J Freshw Ecol. 17:521–529.
Han, M. -S.., Ryu, J. -K.., Yoo, K. -I.. & Kong, D. -S. 1993. Ecological studies on Pal’tang River-Reservoir System in Korea. 1. Annual variation of water quality: past and present. Korean J Limnol. 26:141–149.
Han, M. -S.., Yi, D. -S.., Ryu, J. -K.. & Yoo, K. -I. 1999. Ecological Studies on Pal’tang River-Reservoir System in Korea 3. Photosynthetic parameters and primary productivity of phytoplankton. Korean J Limnol. 32:8–15.
Hall, TA. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symp Ser. 41:95–98.
Ichimura, T. & Itoh, T. 1977. Preservation methods of microalgae: preservation methods of microorganisms. University of Tokyo Press, Tokyo, 355–373, (in Japanese).
Johansen, JR., Bohunická, M., Lukeşová, A., Hrčhková, K., Vaccarino, MA. & Chesarino, NM. 2014. Morphological and molecular characterization within 26 strains of the genus Cylindrospermum (Nostocaceae, Cyanobacteria), with descriptions of three new species. J Phycol. 50:187–202.
Joo, J. -H.., Park, BS., Lee, E. -S.., Kang, Y. -H.. & Han, M. -S. 2016a. Inhibition of growth and microcystin toxicity, and characterization of algicidal substances from Lactobacillus graminis against Microcystis aeruginosa
. Korean J Ecol Environ. 49:176–186.
Joo, J. -H.., Park, C. -S.., Choi, HJ., Lee, HW. & Han, M-S. 2017. A field application feasibility of biologically derived substances (naphthoquinone derivate: NQ 2-0) for the mitigation of harmful cyanobacterial blooms. Ecol Resil Infrastruct. 4:130–141.
Joo, J. -H.., Park, C. -S.., Kuang, Z., Byun, J. -H.., Lee, HW., Choi, HJ. & Han, M-S. 2016b. Development of immobilized naphthoquinone for effective algicidal activity under various environmental conditions and it’s ecological changing monitoring. Korean J Environ Biol. 34:281–291.
Kim, BH., Lee, WS., Kim, Y. -O.., Lee, H. -O.. & Han, M-S. 2005. Relationship between akinete germination and vegetative population of Anabaena flos-aquae (Nostocales, Cyanobacteria) in Seokchon reservoir (Seoul, Korea). Arch Hydrobiol. 163:49–64.
Komárek, J. & Mareş, J. 2012. An update to modern taxonomy (2011) of freshwater planktic heterocytous cyanobacteria. Hydrobiologia. 698:327–351.
Laloui, W., Palinska, KA., Rippka, R., Partensky, F., de Marsac, NT., Herdman, M. & Iteman, I. 2002. Genotyping of axenic and non-axenic isolates of the genus Prochlorococcus and the OMF-’Synechococcus’ clade by size, sequence analysis or RFLP of the internal transcribed spacer of the ribosomal operon. Microbiology. 148:453–465.
Lee, D. -S.., Suh, S. -O.., Hwang, S. -K.., Kwon, T. -K., Kim, T. -H.., Shin, W. -C.. & Hong, S-D. 1996. Nucleotide sequence of 16S rRNA gene from Streptomyces melanosporofaciens 7489. J Microbiol Biotechnol. 6:364–365.
Li, Z. & Brand, J. 2007.
Leptolyngbya nodulosa sp. nov. (Oscillatoriaceae), a subtropical marine cyanobacterium that produces a unique multicellular structure. Phycologia. 46:396–401.
Li, Z., Han, M. -S.., Hwang, S. -O.., Byeon, M. -S.., Hwang, S. -J.. & Kim, B-H. 2013. Molecular identification of the bloom-forming cyanobacterium Anabaena from north Han River system in summer 2012. Korean J Ecol Environ. 46:154–162.
Li, Z., Shin, HH., Lee, T. & Han, M-S. 2015. Resting stages of freshwater algae from surface sediments in Paldang Dam Lake, Korea. Nova Hedwigia. 101:475–500.
Ludwig, W., Strunk, O., Klugbauer, S., Klugbauer, N., Weizenegger, M., Neumaier, J., Bachleitner, M. & Schleifer, KH. 1998. Bacterial phylogeny based on comparative sequence analysis (review). Electrophoresis. 19:554–568.
Lukešová, A., Johansen, JR., Martin, MP. & Casamatta, DA. 2009.
Aulosira bohemensis sp. nov.: further phylogenetic uncertainty at the base of the Nostocales (Cyanobacteria). Phycologia. 48:118–129.
Mishra, S., Bhargava, P., Adhikary, SP., Pradeep, A. & Rai, LC. 2015. Weighted morphology: a new approach towards phylogenetic assessment of Nostocales (Cyanobacteria). Protoplasma. 252:145–163.
Nübel, U., Garcia-Pichel, F. & Muyzer, G. 1997. PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl Environ Microbiol. 63:3327–3332.
Nylander, JA. 2004. MrModeltest ver. 2.3 Computer software and manual. Evolutionary Biology Centre, Uppsala University, Uppsala.
Oliver, RL. & Ganf, GG. 2000. Freshwater blooms. In : Whitton B, Potts M, editors The Ecology of Cyanobacteria: Their Diversity in Time and Space. Kluwer Academic, Dordrecht, 149–194.
Pearson, L., Mihali, T., Moffitt, M., Kellmann, R. & Neilan, B. 2010. On the chemistry, toxicology and genetics of the cyanobacterial toxins, microcystin, nodularin, saxitoxin and cylindrospermopsin. Mar Drugs. 8:1650–1680.
Rajaniemi, P., Hrouzek, P., Kastoyská, K., Willame, R., Rantala, A., Hoffmann, L., Komárek, J. & Sivonen, K. 2005. Phylogenetic and morphological evaluation of the genera Anabaena, Aphanizomenon, Trichormus and Nostoc (Nostocales, Cyanobacteria). Int J Syst Evol Microbiol. 55:11–26.
Rippka, R., Deruelles, J., Waterbury, JB., Herdman, M. & Stanier, RY. 1979. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology. 111:1–61.
Ronquist, F. & Huelsenbeck, JP. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 19:1572–1574.
Rudi, K., Skulberg, OM. & Jakobsen, KS. 1998. Evolution of cyanobacteria by exchange of genetic material among phyletically related strains. J Bacteriol. 180:3453–3461.
Singh, P., Fatma, A. & Mishra, AK. 2015. Molecular phylogeny and evogenomics of heterocytous cyanobacteria using rbcL gene sequence data. Ann Microbiol. 65:799–807.
Stackebrandt, E. & Goebel, BM. 1994. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol. 44:846–849.
Suh, M. -Y.., Kim, B. -H.., Bae, K. -S.. & Han, M-S. 2005. Annual fluctuation (2000–2003) of water quality and cyanobacterial abundance in the lower part of Han-River. Korean J Limnol. 38:181–187.
Tanabe, Y., Kasai, F. & Watanabe, MM. 2007. Multilocus sequence typing (MLST) reveals high genetic diversity and clonal population structure of the toxic cyanobacterium Microcystis aeruginosa
. Microbiology. 153:3695–3703.
Tuji, A. & Niiyama, Y. 2010. Phylogenetic study by the morphological and molecular analyses of Japanese planktonic Anabaena species. Bull Natl Mus Nat Sci Ser B. 36:71–80.
Urbach, E., Robertson, DL. & Chisholm, SW. 1992. Multiple evolutionary origins of prochlorophytes within the cyanobacterial radiation. Nature. 355:267–270.
Vaccarino, MA. & Johansen, JR. 2012.
Brasilonema angustatum sp. nov. (Nostocales), a new filamentous cyanobacterial species from the Hawaiian islands. J Phycol. 48:1178–1186.
Wacklin, P., Hoffmann, L. & Komárek, J. 2009. Nomenclatural validation of the genetically revised cyanobacterial genus Dolichospermum (Ralfs ex Bornet et Flahault) comb. nova. Fottea. 9:59–64.
Watanabe, M., Niiyama, Y. & Tuji, A. 2004. Studies on planktonic blue-green algae 10. Classification of planktonic Anabaena with coiled trichomes maintained in the national science museum, Tokyo. Bull Natl Sci Mus Ser B Bot. 30:135–149.
Wilmotte, A. & Herdman, M. 2001. Phylogenetic relationships among the cyanobacteria based on 16S rRNA sequences. In : Garrity GM, Boone DR, Castenholz RW, editors Bergey’s Manual of Systematic Bacteriology. 2nd ed. Vol. 1. The Archaea and the Deeply Branching and Phototrophic Bacteria. Springer, New York, 487–493.
Zapomělová, E., Hisem, D., Řeháková, K., Hrouzek, P., Jezberová, J., Komárková, J., Korelusová, J. & Znachor, P. 2008. Experimental comparison of phenotypical plasticity and growth demands of two strains from the Anabaena circinalis/A. crassa complex (Cyanobacteria). J Plankton Res. 30:1257–1269.
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||